Authors: Sarah Walsh1,2 and Tom Brown2
1. ATDBio Ltd, Oxford Science Park, Oxford OX4 4GA, UK.
2. Chemistry Research Laboratory, University of Oxford, Oxford OX1 3TA, UK.
Fluorescence is used extensively in nucleic acid-based applications such as DNA sequencing, real-time PCR and genome imaging.1,2 Well-established fluorescence systems for nucleic acid sequence recognition include TaqMan probes,3 Molecular Beacons,4 Scorpion primers5 and HyBeacons.6 In all these cases, a fluorophore is combined with a quencher (fluorescent or non-fluorescent) to allow fluorescence enhancement in the presence of the target sequence via separation of the fluorophore from the quencher. Improvements in fluorogenic hybridisation probe methodologies have great potential in the field of nucleic acid-based diagnostics, and in this context, thiazole orange has been the focus of intense study.
Thiazole orange is a fluorescent asymmetric cyanine dye with an excitation peak at 514 nm and an emission peak at 533 nm. The molecule can be excited using a 488 nm laser and is composed of two heterocyclic ring systems (quinoline and benzothiazole) connected through a methine bridge (Figure 1). The fluorescence intensity of TO depends upon its conformation.7 A planar state allows conjugation between the two aromatic systems- this is the fluorescent form; whereas rotation at the methine bridge produces a non-planar conformation, which is not fluorescent. In the presence of double stranded (ds) DNA, TO acts as an intercalator (or groove binder).8,9 When intercalated, its fluorescent planar conformation is stabilised by stacking between base pairs.10 This has led to the use of TO and its analogues such as SyBr Green and TOTO (Figure 1) in the fluorescence detection of dsDNA.11 These molecules are essentially indiscriminate dsDNA binders.
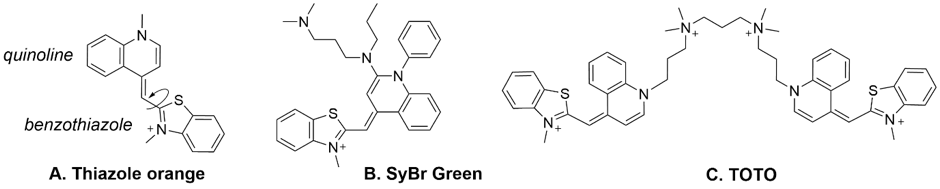
Figure 1: Structure of thiazole orange and other common analogues. The arrow indicates rotation around the methine bridge and only the planar state is fluorescent
In order to provide sequence-specific recognition of target nucleic acids, TO can be attached to oligonucleotide probes. Such TO oligonucleotide conjugates have been used in combination with other fluorophores for highly sensitive, multi-colour detection of DNA and RNA targets.12,13 The TO moiety has the useful additional property of strongly stabilising DNA duplexes, thus allowing shorter probes to be used and increasing discrimination between wild-type and mutant target sequences.13 Pioneering work has been carried out by the Seitz group (FIT probes) using TO-labelled PNA14 and TO-labelled oligonucleotides15 for applications that include live cell studies.16 The contributions to the field by Okamoto (ECHO probes)17,18 and Wagenknecht (artificial TO DNA base)19 are also ground-breaking. Furthermore, TO has been tethered to triplex forming oligonucleotides (TFOs) for sequence specific detection of DNA duplexes.20 This approach provides remarkable triplex stabilisation and expands the potential range of duplex targets to include base pair inversions at biologically relevant pH. With these TO-TFOs, the stabilising effects of TO are additive, with the most stable triplex at neutral pH evidenced by a ∆Tm = +45 °C (compared to that of the unmodified TFO). This stability is accompanied by large enhancements in fluorescence (26-fold increase at pH 7) and quantum yields (up to 40-fold).20
The simplest way to incorporate TO into an oligonucleotide is to employ its NHS ester in post synthetic labelling of the corresponding amino-modified oligonucleotide. Various amino-modifiers are commercially available for incorporation into oligonucleotides at the termini or internally, and the labelling method is straightforward and generally high yielding. One example is the use of an internal amino-modifier C6 dT nucleobase (Figure 2A), which places the fluorescent label in the major groove upon duplex formation. Once the oligonucleotides are labelled, purification is carried out via HPLC (Figure 2B).
| A. |
|
|
| B. |
|
Figure 2. Post-synthetic labelling of oligonucleotides with TO. A. Labelling of internal thymine bases using TO NHS ester (in box) and amino-modifier C6 dT. B. UPLC analysis of purified oligonucleotides labelled with multiple TO units.20 Retention time increases significantly as a function of the number of TO additions. |