Due to their sequence specific targeting nature, oligonucleotides have long been considered valuable tools in drug development. This is because once a therapeutic platform is fully developed, it can theoretically be applied to any target/disease by simply changing the oligonucleotide sequence. To date, platforms have included antisense and RNA interference, which have given rise to close to ten FDA-approved drugs.
Examining the sequences of these oligonucleotide drugs, reveals many different modifications to the sugar-phosphate backbone.1, 2 These modifications 1) modulate the affinity of oligonucleotide drugs to their complementary targets, and/or 2) improve the stability/half-life of pharmaceutically active compound in cells and human fluids. Modified oligonucleotides are synthesized on the same synthesizers used for unmodified sequences and require special reagents, many of which are available at Glen Research. In addition to synthesis reagents for phosphorothioates (PS), 2'-OMe RNA ( 2'-OMe) and 2’-F RNA, Glen Research also offers other attractive modifications such as LNA, DNA PACE, 2'-OMe-RNA PACE, methyl phosphonates, 2’-FANA, DNA phosphorodithioates, 2'-OMe RNA phosphorodithioates and L-DNA, which can all be utilized in the development of oligonucleotide therapies to varying degrees.
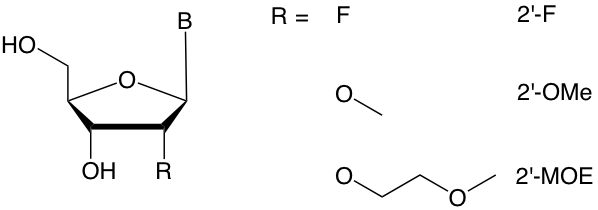
Figure 1. RNA 2’-modified backbones
A backbone modification that has been particularly prominent is 2’-O-methoxyethyl-RNA (2’-MOE).3 Like other 2’ modifications of ribose nucleotides (Figure 1), 2’-MOE favors the formation of A-form, RNA-like double helices, resulting in enhanced duplex stability when paired with RNA targets. In addition, 2’-MOE provides significant nuclease resistance and is relatively non-toxic. This combination has made 2’-MOE an attractive backbone for many oligonucleotide drug candidates, three of which have been approved by the FDA (Table 1 and Figure 2).4, 5
|
Drug |
Condition |
Mechanism |
FDA Approval |
|
Kynamro (Mipomirsen) |
Familial hypercholestolemia |
RNase H |
2013 |
|
Spinraza (Nusinersen) |
Spinal muscular atrophy |
Splicing modulation |
2016 |
|
Tegsedi (Inotersen) |
Hereditary transthyretin-mediated amyloidosis |
RNase H |
2018 |
Table 1. 2’-MOE oligonucleotide drugs approved by the FDA
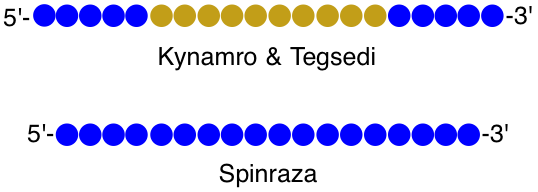
Figure 2. Structure schemes of three 2’-MOE drugs. Blue denotes 2’-MOE; tan denotes DNA; all phosphates are PS.
Kynamro treats homozygous familial hypercholesterolemia, a rare cholesterol disease, while Tegsedi treats nerve damage associated with hereditary transthyretin-mediated amyloidosis. Both drugs are 2’-MOE/DNA chimeras (gapmers) that contain ten central DNA nucleotides and five 2’-MOE nucleotides at each terminus. The two 2’-MOE regions enhance nuclease stability while ensuring that the oligonucleotide binds to its complementary sequence with appropriate affinity. The central DNA region, upon binding to target mRNA, forms a DNA/RNA heteroduplex, which is a substrate for RNase H cleavage. For Kynamro, mRNA cleavage results in a reduction of apolipoprotein B-100, a major component of low-density and very low-density lipoproteins. For Tegsedi, circulating concentrations of transthyretin are significantly reduced.
Spinraza was the first drug approved for spinal muscular atrophy, a rare disease that is the most common genetic cause of infant deaths. It is an eighteen nucleotide all 2’-MOE sequence, and unlike Kynamro and Tegsedi, Spinraza is a steric blocking oligonucleotide. When target mRNA is bound, no cleavage occurs. Instead, the binding of Spinraza onto its target mRNA interferes with splicing mechanisms, allowing an extra exon to be retained and rescuing the production of functional survival motor neuron 1 (SMN1) protein. When treated early with Spinraza, patients have much better outcomes, including superior motor coordination and reduced death rates.
In terms of balancing toxicity, affinity/off-target effects, and nuclease stability, the 2’-MOE backbone plays an important role in all three of the aforementioned drugs. As research with 2’- MOE continues to be conducted, Glen Research has decided to make the backbone more accessible by adding the 2’-MOE phosphoramidites of A, 5-Me-C, G and 5-Me-U (Figure 3).
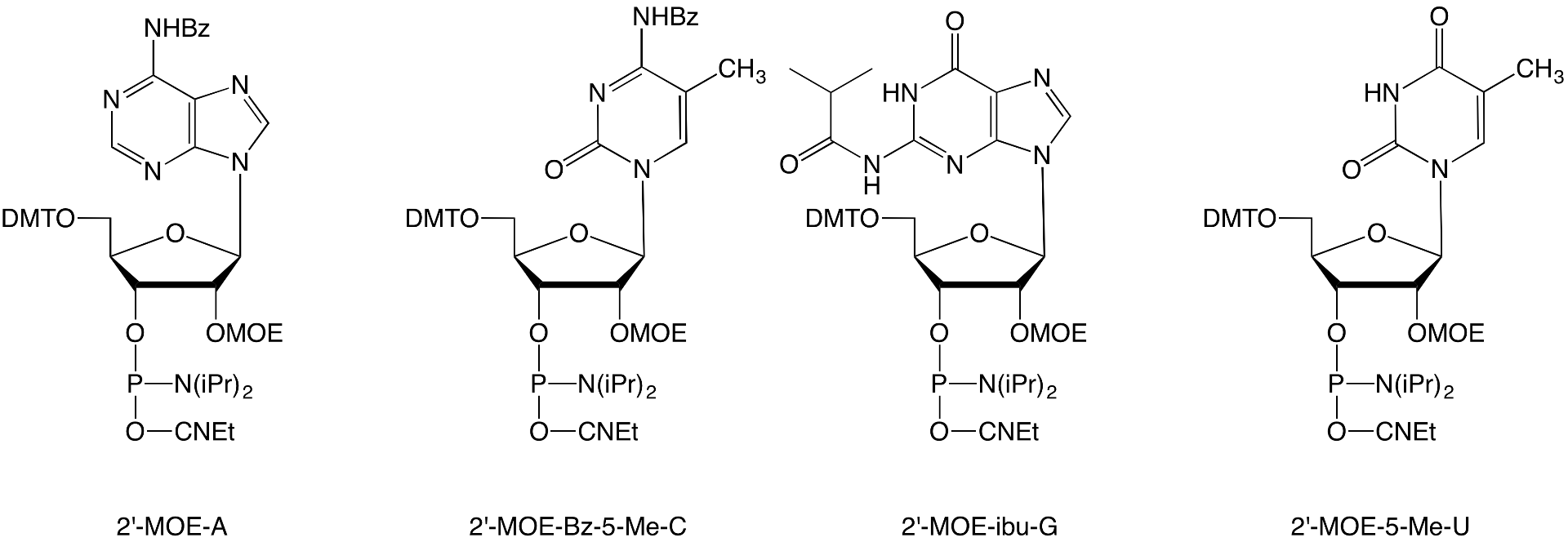
Figure 3. 2’-MOE Phosphoramidites
Although 2’-MOE and 2'-OMe are very similar in terms of chemistry and functionality, 2'-OMe has been used in a much broader range of applications, some of which are unrelated to therapeutics. These include aptamers,6 detection probes,7 RNAi,8 DNAzymes/ribozymes9, 10 and CRISPR.11 2’-MOE should be applicable in all these contexts, as it has superior duplex stability and nuclease resistance. We hope that the addition of 2’-MOE will enable researchers to expand in their use of it.
The use of 2’-MOE reagents in oligonucleotide synthesis is relatively straightforward. A coupling time of 6 min is recommended, and oligonucleotides that contain these residues can be deprotected following our standard procedures. It is important to note that methylamine should not be used with 2'-MOE-Bz-5-Me-C, in order to avoid methylation of the N4.