Figure 1: structures of Ribonucleosides Supported by Glen Research
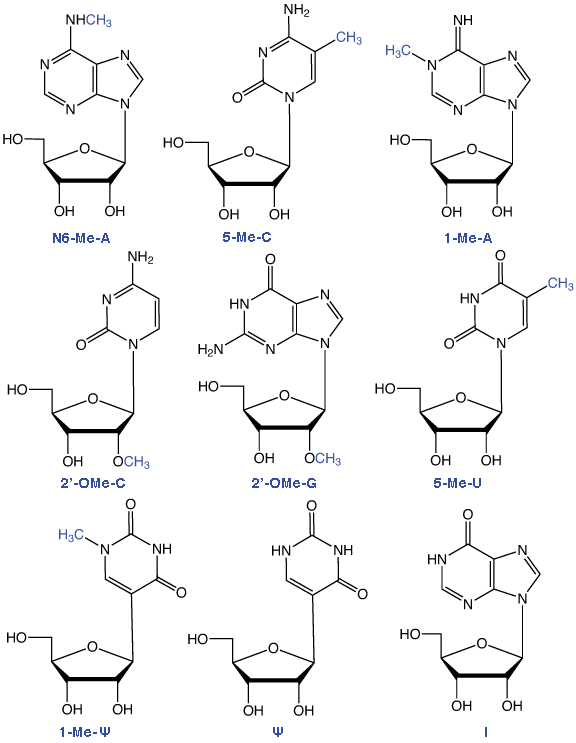
RNA methylation occurs in a large selection of RNA nucleosides and this post transcriptional modification of RNA, carried out by a variety of RNA methyltransferases, appears in a wide variety of RNA species - including tRNA, but also mRNA, miRNA and RNA viruses. The most common methylated ribonucleoside in eukaryotes is N6-methyladenosine (m6A). In an earlier Glen Report 26.2, page 1, Qing Dai and Chuan He of the University of Chicago discussed the recent finding that the behavior of m6A offers an example of reversible RNA methylation with a significant role as an epigenetic modification.1 In this article, we will focus on the role of methylated ribonucleosides in tRNA and mRNA.
A recent review2 reveals that over 90 methylated nucleosides have been found in tRNA and that these play a significant role in the stabilization of tRNA structure, reinforcement of the codon-anticodon interaction, regulation of wobble base pairing, and prevention of frameshift errors. In addition, methylation appears to mark the tRNA as mature, preventing its degradation as well as directing localization within the cell.
Ever since it was discovered3 that injecting mRNA directly into mouse skeletal muscle led to the expression of the encoded protein, there has been great interest in using mRNA as a therapeutic species. However, the cell's innate immune response, designed to detect the foreign RNA of viruses, can be activated after mRNA delivery, leading to the shutdown of protein translation and ultimately, cell death. It was found, however, that some nucleobase modifications can greatly enhance the properties of mRNA by reducing immunogenicity and increasing stability. Specifically, mRNA modified with pseudouridine (Ψ) alone, or in combination with 5-methylcytidine (5-Me-C), significantly increased translation levels. When Ψ was substituted with 1-methylpseudouridine (1-Me-Ψ), even higher levels of protein expression were observed.4
Further studies showed5 that mRNA containing 1-Me-Ψ alone, or in combination with 5-Me-C, outperformed the equivalent Ψ and/or 5-Me-C mRNA by providing higher reporter gene expression upon transfection into cell lines and mice. It was shown that 1-Me-Ψ/5-Me-C modified mRNA resulted in reduced intracellular innate immunogenicity and improved cellular viability upon in vitro transfection.5
We take this opportunity to introduce 1-Me-Ψ to our catalog of RNA minor bases. 1-Me-Pseudouridine Phosphoramidite is fully compatible with standard RNA synthesis conditions. Similarly, with no protecting groups to be removed, oligo deprotection can be carried out under standard conditions, including our preferred deprotection with AMA at 65°C for 10 minutes.
All of the RNA minor bases shown in Figure 1 are available from Glen Research as phosphoramidites. In future, we will continue our introduction of minor bases involved in RNA epigenetics and methylated RNA in the hope and expectation that their availability as phosphoramidites will act as a catalyst to promote these exciting research activities.
Figure 2: Structure of 1-Me-Pseudouridine |
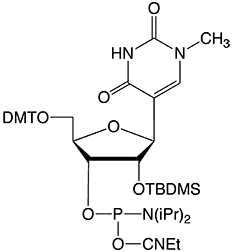 |
1-Me-Pseudouridine Phosphoramidite |
1. Y. Fu, D. Dominissini, G. Rechavi, and C. He, Nat Rev Genet, 2014, 15, 293-306.
2. H. Hori, Front Genet, 2014, 5, 144.
3. J.A. Wolff, et al., Science, 1990, 247, 1465-8.
4. D. Weissman, Expert Rev Vaccines, 2015, 14, 265-81.
5. O. Andries, et al., J Control Release, 2015, 217, 337-44.