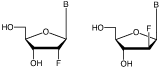
Arabinonucleosides are epimers of ribonucleosides with the chiral switch being at the 2’ position of the sugar residue. In ribonucleosides, the 2’-hydroxyl group is in the bottom face (a) position and, in arabinonucleosides, the 2’-hydroxyl group is in the top face (b) position. The 2’-F versions of the nucleosides in RNA and ANA are shown in Figure 1.
2’-F RNA monomers have been available commercially for several years and oligonucleotides containing this modification have been very useful in antisense, siRNA and aptamer studies. Thanks to the pioneering work of Masad Damha and his group at the Department of Chemistry, McGill University in Montreal, 2’-F-ANA is now attracting considerable attention in the research community. In this article, we briefly review the properties of 2’-F-ANA and assess its promise in these fields of research.
Modification at the 2’ position of nucleosides has proved to be critical in adjusting the properties of oligonucleotides. It is also relatively straightforward to predict the effect of 2’ modification on the conformational geometry of the sugar and its subsequent effect on duplex stability. For example, the electronegative fluorine in 2’-F-RNA causes the sugar to adopt the C3’-endo configuration, normally observed in RNA, which leads to an increase in duplex stability, while conformationally restricted analogues like LNA are fully constrained to that configuration. This leads to a significant increase in melting temperature for each position modified by 2’-F-RNA (DTm 3.0 °C/mod) and LNA (DTm 5.6° C/mod). In contrast, the 2’-F-ANA adopts a more DNA-like B-type helix conformation, not through the typical C2'-endo conformation but, rather, through an unusual O4’-endo (east) pucker. However, the presence of the electronegative fluorine leads to a still significant increase (DTm1.2° C/mod) in melting temperature per modification.1
High base pairing specificity is essential in such applications as diagnostics and therapeutics. 2’-F-ANA-containing oligonucleotides exhibit very high binding specificity to their targets. Indeed, a single mismatch in a 2’-F-ANA – RNA duplex leads to a DTm of -7.2 °C and in a 2’-F-ANA-DNA duplex a DTm of -3.9 °C.2
While adopting a DNA-like structure, 2’-F-ANA is much more stable to enzymatic and chemical hydrolysis. Indeed, the order of stability to snake venom phosphodiesterase has been reported to be phosphorothioate (PS) DNA >> 2’-F-ANA > RNA > 2’-F-RNA, while phosphodiester (PO) 2'-F-ANA > PO-DNA. The glycosidic bond in 2’-F-ANA is also remarkably stable to acidic hydrolysis due to the presence of the electronegative fluorine and 2'-F-ANA is stable at pH 1.2 at 37 °C for several days without cleavage. The presence of fluorine at the 2’ position in 2’-F-ANA also leads to increased stability to hydrolysis under basic conditions relative to RNA and even 2’-F-RNA.1,3 The stability of 2'-F-ANA to nucleases makes this a useful modification for enhancing the stability of oligonucleotides in biological environments.2
2’-F-ANA hybridizes strongly to target RNA and, unlike most 2’ modifications, induces cleavage of the target by RNase H. PS 2’-F-ANA is routinely used in these applications due to its increased nuclease resistance. However, uniformly modified PS 2’-F-ANA, while still active, is not as well recognized by RNase H. Of course, the gapmer strategy of flanking a DNA core in the oligo with PS 2’-F-ANA segments is effective and unusually so. In this case, even a single deoxynucleoside in the gap restores high RNase H activity. Interestingly, the target RNA is cleaved throughout the chain rather than only at positions within the gap.2 Furthermore, alternating 2'-F-ANA and DNA units provide among the highest potency RNase H-activating oligomers. Both the “altimer” and “gapmer” strand architectures consistently outperform PS-DNA and DNA/RNA gapmers.4
Since 2’-F-ANA exhibits mostly DNA-like behavior and siRNA activity requires RNA-like behavior, is there a place for 2’-F-ANA in siRNA development? siRNA is recognized in mammalian cells by the multiprotein RNA-induced silencing complex (RISC) containing the exonuclease Argonaute2. When siRNA is introduced, the sense or passenger strand is removed, while the guide strand, antisense to the target mRNA, is incorporated into the RISC. Interestingly, siRNA oligos were found to tolerate the presence of 2’-F-ANA linkages very well. The whole sense strand can, in fact, be all 2’-F-ANA but potency is improved by incorporating 5 RNA residues towards the 3’ terminus. This increased potency may be attributed to a slight destabilization of the duplex at the 3' sense terminus, which favors loading of the antisense strand into the RISC complex.5 A small number of 2’-F-ANA are well tolerated in the antisense strand but modified antisense strands should be 5’-phosphorylated for proper potency. Also, modification of the 3’ overhang of the antisense strand with 2’-F-ANA leads to increased potency. As can be expected, the serum half life of 2’-F-ANA modified siRNA is also significantly improved relative to unmodified siRNA.2,5
A recent publication5 describes high potency gene silencing with siRNA chimeras containing 2’-F-RNA and/or LNA and 2’-F-ANA. The high efficacy of these chimeras was attributed to the combination of the rigid RNA-like properties of 2’-F-RNA and LNA with the DNA-like properties of 2’-F-ANA. There is clearly great potential for 2’-F-ANA to contribute to the continuing improvement of the efficacy of siRNA.
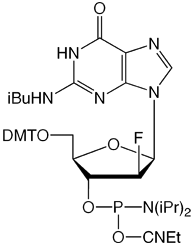
We are happy to introduce all four 2’-F-ANA monomers, as shown in Figure 2. 2’-F-ANA oligonucleotides are readily prepared with a 6 minute coupling time under otherwise standard conditions. Sulfurization, if desired, can be carried out using either of our Sulfurizing Reagents. The resulting oligonucleotides can be deprotected using ammonium hydroxide even at elevated temperatures. Oligonucleotides and chimeras can be purified using all standard methods.



We thank Masad Damha for taking the time to review this article and for his encouragement to make these products available to the research community.
2'-F-ANA is covered by intellectual property. Key patents covering siRNA and antisense applications are as follows:
WO/2009/146556 (siRNA); WO 03064441 and WO 0220773 (antisense).