Since the beginning of automatic oligonucleotide synthesis in the late 80’s, the possibilities available to researchers have expanded exponentially. Starting from simple primers for sequencing, PCR or cloning, oligos are now used for gene expression modulation1, as probes for quantitative PCR2,3,4, and even for some futuristic applications in nanotechnology such as logic-gates5.
Controlling DNA functions using an external stimulus that could be easily and quantitatively controlled at any given moment and position might be an incredible and interesting way to develop novel applications in cell biology and pharmacology. It could also provide new tools for the analysis of the mechanisms underlying DNA recognition and DNA-mediated bioprocesses.
Photo-control, the use of ultraviolet or visible light to control a reaction, has a number of advantages over other external stimuli:
When a photo-responsive molecule is directly attached to DNA as a receptor, photo-regulation of the bioprocess regulated by that DNA molecule could, in principle, be achieved. Such photo-responsive DNA could also be used as a switch in a DNA-based nano-machine.
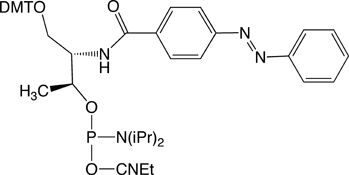
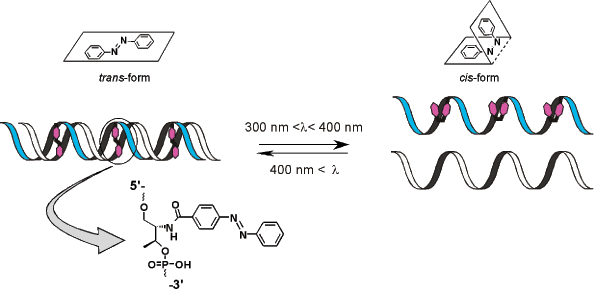
Professor Hiroyuki Asanuma and his group at the department of Molecular Design and Engineering of the Graduate School of Engineering of the Nagoya University (Japan) have developed an efficient method to achieve this goal. They have attached azobenzene to DNA and made it photo-responsive6,7. Azobenzene is a typical photo-responsive molecule that isomerizes from its planar trans-form to the non-planar cis-form after UV-light irradiation with a wavelength between 300 nm and 400 nm (lmax is around 330 nm). Interestingly, the system reverts from the cis-form to the trans-form after further irradiation with visible light (wavelength over 400 nm). This process is completely reversible, and the azobenzene group does not decompose or induce undesirable side reactions even on repeated trans-cis isomerization.
By introducing azobenzenes into DNA through D-threoninol as a linker, Asanuma and co-workers succeeded in achieving photo-regulation of:
For photo-regulation of DNA hybridization or DNA transcription as effected by T7-RNA polymerase, the azobenzene [X] residues must be inserted between the base pairs of the template DNA duplex. In other words, the X residue is introduced as a bulge adduct. For example, to make the sequence 5’-GCGAGTCG-3’ photo-responsive, the X residue must be incorporated in the sequence, without any base replacement to obtain 5’-GCGAXGTCG-3’. This sequence’s complementary strand, 3’-CGCTCAGC-5’, remains composed only of natural nucleotides. As long as the X residue is additionally inserted, there are no limitations with regard to the position of its introduction. X can equally be inserted at the center of a sequence or in the vicinity of its 5’- or 3’-terminus.
The introduction of several X residues increases the DTm created by the photo-induced isomerization. For example, with the sequence described in the previous paragraph with one X residue in the middle, the DTm between the trans [more stable] form and the cis form is 14.3°C and with 2 residues as in 5’-G CXG AGT XCG-3’ the DTm between cis and trans is 21.5°C.
In order to photo-regulate the hybridization of relatively long duplexes efficiently, introduction of multiple azobenzenes is recommended. In this case as well, X residues should be additionally introduced without replacing any bases. A photo-responsive DNA sequence is designed as 5’-(NNX)n-NN-3’. If chemical modification is allowed to both strands, azobenzenes can be symmetrically introduced to both strands (i.e., 5’-(NNX)n-NN-3’/3’-(NXNn-XN-5’) to further raise the photoregulatory efficiency13.
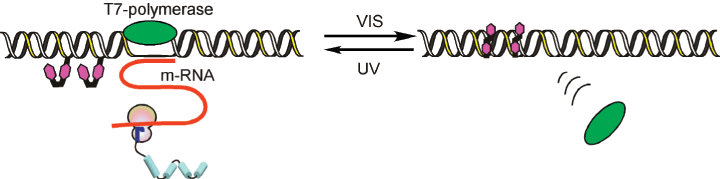
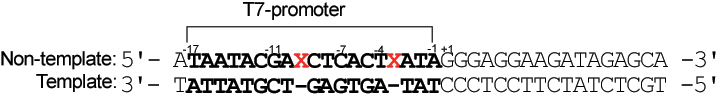
This application was described in detail in a Nature Protocols paper12. In brief, adding azobenzene groups to the T7 promoter allows transcription by T7-RNA polymerase to be reversibly photo-regulated by the trans–cis isomerization of the labels. Transcription is suppressed by visible-light irradiation, which triggers the formation of trans-azobenzene, and is recovered by UV-light irradiation (formation of cis-azobenzene), as shown in Figure 2. For effective photo-regulation, two X residues are inserted between –3 and –4 and between –9 and –10 in the non-template strand of the T7 promoter region (see Figure 3 From Nature Protocols12 - erratum). The template strand is kept intact.
Introduction of the X labeled promoter into gene construct for effective photo-regulation of gene expression in vitro was described in Liang, et al.14
Oligonucleotides containing azobenzene modifiers have also been used in order to control the activity of some enzymes. Tan’s group15 has linked an azobenzene-containing sequence to a Thrombin-binding aptamer via a Spacer-18 residue. This molecule contains 3 parts: an inhibitory part, that is the Thrombin-aptamer, a regulatory part, that is a complementary oligo designed to bind to the aptamer part and contains azobenzenes and a regulatory part, and a linker – in this case Spacer 18. Depending on light exposure, the regulatory part will hybridize to the aptamer – blocking the inhibition – or not – releasing the inhibitory activity.
Using the properties of the photoisomerization, they have been able to optimize a probe whose inhibitory effect can be inhibited or restored using photo-exposition. Possible applications of the technology are also described in this paper.
The synthesis of azobenzene-containing oligos can be performed without any modification using a coupling time of 10 minutes for the coupling of Azobenzene Phosphoramidite. The oligos can be deprotected using standard procedures. The Nature Protocols paper12 described methods for purification of the labelled oligos using HPLC, PAGE or Poly-Pak cartridges.
For the measurement of the concentration by measure of the absorption, the use of an extinction coefficient of 4,100 M-1.cm-1 at 260 nm is recommended.
For UV irradiation, a hand-held, conventional UV illuminator can be used. Although the time required for cis isomerization depends on the intensity of the UV light, 5 min irradiation will usually be enough to achieve a steady state, irrespective of the light source. More information about irradiation using a Xenon-lamp and filter is available in the Nature Protocols paper12.
Oligonucleotides containing azobenzene are easily prepared using a preferred coupling time of 10 minutes. They are deprotected according to the conditions required to deprotect the nucleobases. AMA at 65 °C or UltraMild chemistry can be used. However, deprotection with ammonium hydroxide has to be carried out at room temperature to avoid degradation. For example, no degradation was observed after 17 hours at room temperature using ammonium hydroxide.
Obviously, cis and trans isomers exist for azobenzene analogues, which may complicate HPLC analysis of the monomer and the oligos produced. Using a simple oligonucleotide, 5'-azobenzene-T6, we illustrate the potential for isomerism. In Figure 4, the HPLC shows two peaks for the simple 5'-azobenzene-T6, the earlier eluting and minor peak being the cis isomer and the later eluting peak being the trans isomer. The first peak in the chromatogram is a T6 standard added as the internal standard. On irradiation at 330nm, the ratio changes as the trans isomer converts to the cis isomer. Figure 5 shows the UV spectra of the pure cis and trans isomers.



We are happy to introduce this new and interesting product to our repertoire of modifiers. We would also like to thank Professor Asanuma for his help in preparing this article. In Japan, this product is only available through our distributor Nihon Techno Service Co., Ltd (http://www.ntsbio.com/).