Glen Report 9.14: 3'-Labelling - Dabcyl CPG, Photolabile 3'-Amino-CPG
Dabcyl CPG
The hybridization of an oligonucleotide to its complementary strand is one of the most specific molecular recognition events. Consequently, labelled oligonucleotides have become incredibly powerful tools as diagnostic probes for the determination of gene function as well as for the detection of diseases. One drawback in the process is the fact that non-hybridized probe which would lead to background signal must be removed prior to the detection event. A recent article describes1 the general structure of probes which have the potential to allow real-time measurement. The authors describe these probes as molecular beacons.
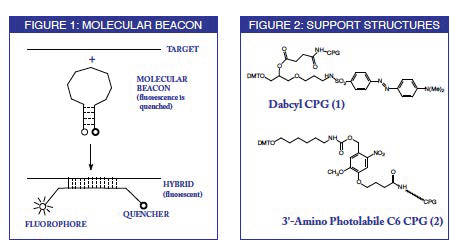
A molecular beacon probe has its natural fluorescence quenched in solution unless it is hybridized to the target sequence. Consequently, the design of a molecular beacon requires a fluorophore to be at one terminus and a quencher molecule to be at the other, with both molecules being separated from the oligonucleotide by a hydrocarbon spacer. The probe sequence is in the center of the molecule and the bases towards both termini are self-complementary and about 5 - 8 nucleotides in length. The lengths of the probe sequences are chosen to maximize the separation of the fluorophore and quencher molecules when the probe is hybridized to the target as illustrated in Figure 1. This occurs when the probe sequence is 15, 25 or 35 nucleotides in length, where the open arms of the molecular beacon are arrayed in a trans configuration.
A number of fluorophore quencher sets could be easily postulated but the chemical synthesis of these doubly-labelled oligonucleotides may be arduous. The synthesis can be accomplished by conjugating an amine-specific fluorophore and a thiol-specific quencher (or vice versa) with the suitably modified oligonucleotide. A simpler approach would involve the use of a universal quencher support for 3'-terminal labelling along with a 5'-fluorophore phosphoramidite. We are offering the dabcyl support (1) as a potential basis for a universal quencher.
Product Information
3'-Amino Photolabile CPG
In late 1995, Glen Research introduced the first commercial photo-cleavable support. This product, 3'-Carboxylate Photolabile C6 CPG, is designed2 predominantly for the synthesis of fully protected oligonucleotide 3'-carboxylic acids. However, it can also be used without photolysis to yield fully deprotected oligonucleotide 3'-carboxylic acids. It is sold under license from Colorado State University. Typically these photolysis reactions are carried out in a photochemical reactor set at the optimum wavelength. However, these photocleavage reactions may also be carried out on a long wavelength (365nm) transilluminator of the type used in most biology labs.
Now, a similar strategy3 allows 3'-alkylamino oligonucleotides to be prepared using 3'-Amino Photolabile C6 CPG (2). After conventional synthesis, oligonucleotides can be cleaved from the support photochemically to release the fully protected 3'-amino oligonucleotides into solution. Conjugation of a hapten to the amino group can then be carried out in organic solution. This avoids the need for the vast excess of hapten which is used routinely in conjugation reactions with fully deprotected oligonucleotides in aqueous solution, typically at pH 9. It also makes removal of excess hapten much more facile.
Product Information
3'-Carboxylate Photolabile C6 CPG and 3'-Amino Photolabile C6 CPG have been discontinued
References
(1) S. Tyagi and F.R. Kramer, Nature Biotechnology, 1996, 14, 303-308.
(2) D.J. Yoo and M.M. Greenberg, J. Org. Chem., 1995, 60, 3358-3364.
(3) D.L. McMinn and M.M. Greenberg, Tetrahedron, 1996, 52, 3827-3840.
- Glen Report 9.11: A Novel Transfection Reagent
- Glen Report 9.12: Protecting Groups for DNA, RNA and 2'-OMe-RNA Monomers
- Glen Report 9.13: A Gallery of Recent Additions
- Glen Report 9.14: 3'-Labelling - Dabcyl CPG, Photolabile 3'-Amino-CPG
- Glen Report 9.15: Properties of Oligonucleotides Containing the Bases P and K
- Glen Report 9.16: Non-Aqueous Oxidation with 10-Camphorsulfonyl-Oxaziridine
- Glen Report 9.17: Cyclic Oligonucleotides
- Glen Report 9.18: DNA Synthesis Columns