Glen Report 23.25: New Product - NPOM-Caged-dT
Photocaging has developed into an attractive and simple procedure for achieving exquisite control over complex biological processes through photoactivation by UV light. The term ‘caging’ refers to the attachment of a photolabile protecting group to a biologically active molecule at specific locations, effectively rendering the molecule inactive. After irradiation by light at the wavelength required to remove the protecting group, the biological properties of the ‘uncaged’ molecule are restored. Glen Research’s interest lies in the caging of nucleoside phosphoramidites used to prepare caged oligonucleotides whose function is restored after uncaging by UV light at a wavelength that causes no DNA damage. In this article, we focus on the approach of Deiters and co-workers at North Carolina State University and introduce a caged monomer to allow our customers to follow their lead.
The particular form of photochemical DNA activation used by the Deiters's group has been detailed in two recent review articles.1,2 In their approach, the nucleobase is caged with the photolabile group, 6-nitropiperonyloxymethyl (NPOM), which can be removed in a few minutes using UV light at 365nm.3 This wavelength and the low intensity of the light used minimizes possible irradiation-induced damage to DNA, which typically occurs at wavelengths below 300nm.4,5
NPOM-Caged dT-CE Phosphoramidite (1) is shown in Figure 1. By attaching the NPOM group to the N3 position of thymidine, the ability of the nucleotide to undergo base-pair formation is inhibited, thus preventing regular hybridization. NPOM-caged dT is the most popular photocaged product to be introduced into DNA to date, although a caged dG monomer has also been described.6
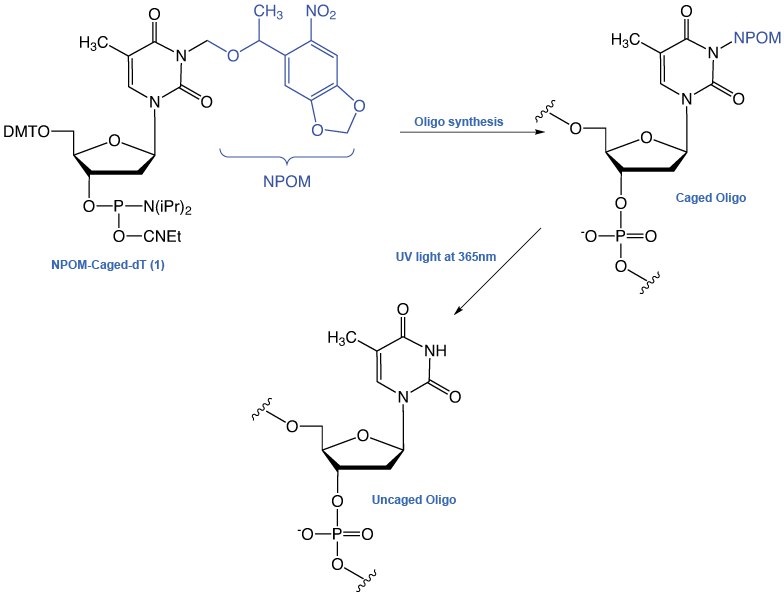
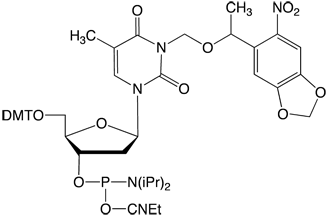
Deiters and coworkers have demonstrated an astonishing array of applications of this strategy in the fields of PCR and polymerase activity7, antisense and gene silencing8, regulation of restriction endonuclease activity9, and enzyme-free mutagenesis10. More recent publications have described activation and deactivation of DNAzyme and antisense activity controlling gene expression in mammalian cells11, and photochemical control of DNA decoy function to regulate gene transcription12.
Oligonucleotides containing NPOM-caged dT are easily prepared by standard oligonucleotide synthesis, with a 10 minute coupling time being optimal for NPOM-caged dT. Cleavage and deprotection of oligonucleotides containing NPOM-caged dT are also carried out under conventional conditions.
As a general design rule, oligonucleotides containing NPOM-caged dT every five or six bases do not hybridize to their complementary strand. However, fewer additions may be tolerated and efficient hybridization was still observed when only a single NPOM-caged dT residue was included.7 Photo-uncaging of an oligonucleotide is easily carried out with UV light at 365 nm for seconds to minutes and this can be readily achieved with a UV transilluminator, a hand-held UV light, or a fluorescence microscope to uncage the oligo in a specific location within the cell.
We thank Professor Deiters for his conversations regarding the potential of caged monomers, for his encouragement to make NPOM-caged dT available to our customers, and for taking the time to review this article.
References
- D.D. Young, and A. Deiters, Org Biomol Chem, 2007, 5, 999-1005.
- A. Deiters, Curr Opin Chem Biol, 2009, 13, 678-86.
- H. Lusic, D.D. Young, M.O. Lively, and A. Deiters, Org Lett, 2007, 9, 1903-6.
- J. Olejnik, E. Krzymanska-Olejnik, and K.J. Rothschild, Nucleic Acids Res, 1996, 24, 361-6.
- J. Cadet, and P. Vigny, The photochemistry of nucleic acids. In Bioorganic Photochemistry, H. Morrison, Ed. John Wiley & Sons: New York, NY, 1990; Vol. 1, pp 179-184.
- H. Lusic, M.O. Lively, and A. Deiters, Mol Biosyst, 2008, 4, 508-11.
- D.D. Young, W.F. Edwards, H. Lusic, M.O. Lively, and A. Deiters, Chem Commun, 2008, 462-4.
- D.D. Young, H. Lusic, M.O. Lively, J.A. Yoder, and A. Deiters, ChemBioChem, 2008, 9, 2937-40.
- D.D. Young, J.M. Govan, M.O. Lively, and A. Deiters, ChemBioChem, 2009, 10, 1612-6.
- D.D. Young, H. Lusic, M.O. Lively, and A. Deiters, Nucleic Acids Res, 2009, 37, e58.
- D.D. Young, M.O. Lively, and A. Deiters, J Am Chem Soc, 2010, 132, 6183-93.
- J.M. Govan, M.O. Lively, and A. Deiters, J Am Chem Soc, 2011, 133, 13176-82.
Ordering Information
- Glen Report 23.21: Glen Research Epigenetic Bases Report
- Glen Report 23.22: New Product – 5’-AminoOxy-Modifier 11
- Glen Report 23.23: New Products - BlackBerry® Quencher (BBQ-650®)
- Glen Report 23.24: On-resin Synthesis of Maleimido-oligonucleotides
- Glen Report 23.25: New Product - NPOM-Caged-dT
- Glen Report 23.26: Glen-Pak™ Purification - Then and Now
- Glen Report 23.27: Technical Brief - Deprotection of HMdU
- Glen Report 23.28:Technical Brief - Improved Oligo Synthesis using 2-Amino-dA and CSO Oxidation
- Glen Report 23.29: New Products - Bulk Polystyrene Supports

