Glen Report 19.17: Selenium Derivatization of Nucleic Acids for X-ray Crystal Structure Determination
Zhen Huang, Ph.D.
Department of Chemistry
Georgia State University
Atlanta, Georgia 30303
Phone: (404) 651-2915 (Office)
Fax: (404) 651-1416
Email: huang@gsu.edu
Lab Homepage: http://chemistry.gsu.edu/faculty/Huang/ (Page no longer available)
Determination of the three-dimensional structures of RNA molecules, RNA-protein and DNA-protein complexes with high resolution is invaluable for gaining understanding of biological systems at the molecular level. X-ray crystallography is the most direct and powerful tool for structure determination of these macromolecules. However, besides the difficulties related to crystallization, heavy atom derivatization for phase determination has been a long-standing problem in nucleic acid X-ray crystallography that has largely slowed down structural determination of new structures and folds. The conventional approaches for DNA and RNA derivatization, such as heavy-atom soaking and co-crystallization, have proved to be much more difficult for nucleic acids than for proteins, probably because nucleic acids often lack specific binding sites for metal ions. In addition, the radiation stability and structure perturbation have reduced the usefulness of the halogen derivatization (such as Br and I).
Recently, we have successfully demonstrated a novel derivatization strategy via selenium replacement of oxygen in nucleic acids.1-8 Unlike conventional halogen derivatization (Br or I), where halogens are primarily placed on the 5-position of deoxyuridine (a mimic of thymidine), selenium can be selectively introduced to a variety of positions via oxygen replacement (e.g., 2’-, 3’-, 5’-ribose oxygen, furan ring oxygen, non-bridging phosphate oxygen, or oxygen on nucleobases).1-8 Choice of positioning can avoid disruption of structure and function caused by modification. We have found that the 2’-Se-derivatized DNA structure (Fig. 1) reveals that the 2’-Se-furanose displays the 3’-endo sugar pucker, which is consistent with the sugar pucker of A-form DNAs and RNAs, and the 2’-methylseleno group is placed in the minor groove of the duplex.
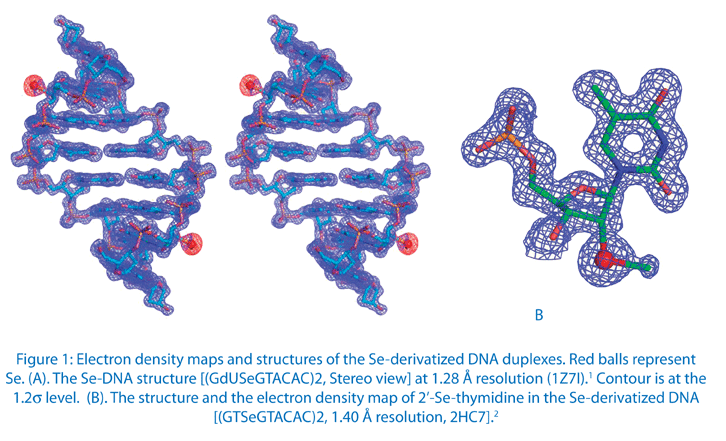
In addition, we found that the local backbone torsion angles and solvent hydration patterns were altered in the DNA structure with the Br derivatization.1 Furthermore, while the native and Br-derivatized DNAs needed over several weeks to form reasonable-size crystals, we observed that the Se-derivatized DNAs grew crystals overnight with high diffraction quality,1,2 suggesting that the Se-derivatization facilitated suggesting that the Se-derivatization facilitated 1,2 suggesting that the Se-derivatization facilitated crystal formation. In addition, the Se-derivatized DNA sequences crystallized under a broader range of buffer conditions, and generally had a faster crystal growth rate. We have observed Se-DNA crystal formation in a few days or even overnight, while the crystals of the corresponding native DNAs need two to three months to grow. Our experimental results indicate that the selenium derivatization of DNAs may largely facilitate the determination of nucleic acid X-ray crystal structures in phasing and high-quality crystal growth. Moreover, our results suggest that the Se derivatization can be a better alternative to the conventional Br derivatization. This Se derivatization strategy via the atom-specific substitution will significantly facilitate X-ray crystal structure studies of nucleic acids and their protein complexes.
References
- Jiansheng Jiang, Jia Sheng, Nicolas Carrasco, and Zhen Huang*, “Selenium Derivatization of Nucleic Acids for Crystallography”, Nucleic Acids Research, 2007, 35, 477-485.
- Jia Sheng, Jiansheng Jiang, Jozef Salon, and Zhen Huang*, “Synthesis of a 2’-Se-Thymidine Phosphoramidite and Its Incorporation into Oligonucleotides for Crystal Structure Study”, Organic Letter, 2007, 9, 749-752.
- Nicolas Carrasco, Julianne Canton-Williams, Gary Brandt, Siming Wang, and Zhen Huang*, “Efficient Enzymatic Synthesis of Phosphoroselenoate RNA Using Adenosine 5’-(?-P-seleno)triphosphate”, Angewandte Chemie Int. Ed., 2006, 45, 94-97.
- Jozef Salon, Guexiong Chen, Yoani Portilla, Markus W. Germann, and Zhen Huang*, “Synthesis of a Novel 2’-Se-uridine Phosphoramidite and Its Incorporation into Oligonucleotides for Structure Study”, Organic Letter, 2005, 7, 5645-5648.
- Nicolas Carrasco, Yuri Buzin, Elizabeth Tyson, Elizer Halpert, and Zhen Huang*, “Selenium derivatization and crystallization of DNA and RNA oligonucleotides for X-ray crystallography using multiple anomalous dispersion”, Nucleic Acids Research, 2004, 32, 1638-1646.
- Nicolas Carrasco and Zhen Huang*, “Enzymatic Synthesis of Phosphoroselenoate DNA using Thymidine? 5’-(? -P-Seleno)-triphosphate and DNA Polymerase for X-ray Crystallography using MAD”, ? Journal of American Chemical Society, 2004, 126, 448-449.
- Quan Du, Nicolas Carrasco, Marlanna Teplova, Christopher J. Wilds, Martin Egli, and Zhen Huang*, “Internal Derivatization of Oligonucleotides with Selenium for X-ray Crystallography Using MAD”, Journal of American Chemical Society., 2002, 124, 24-25.
- Nicolas Carrasco, Dov Ginsburg, Quan Du, and Zhen Huang*, “Synthesis of Selenium-Derivatized Nucleosides and Oligonucleotides for X-ray Crystallography”, Nucleosides, Nucleotides, & Nucleic Acids, 2001, 20, 1723-1734.
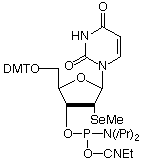
2'-Se-Me-U-CE Phosphoramidite
Product Information
2'-Se-Me-U-CE Phosphoramidite has been discontinued.
- Glen Report 19.11: Modified RNA Phosphoramidites Useful in siRNA Research and Biologically Significant 1-Methyl-adenosine
- Glen Report 19.12: 1-Methyl-Adenine in Nucleic Acids
- Glen Report 19.13: Technical Brief - Precautions During Packaging of Phosphoramidites and Transfer into Alternative Vials
- Glen Report 19.14: Microarrays, Nanotechnology and Beyond
- Glen Report 19.15: 5’-Hexynyl Phosphoramidite – Conjugation with a Click
- Glen Report 19.16: Quenched Autoligation (QUAL) Probes
- Glen Report 19.17: Selenium Derivatization of Nucleic Acids for X-ray Crystal Structure Determination
- Glen Report 19.18: Large Scale Synthesis Update
- Glen Report 19.19: NHS-Carboxy-dT – Expanding the NHS Ester Phosphoramidite Repertoire

