It is well over 20 years since the first amino-modifier prototypes were described on a modified base1 and at the 5’ terminus2. However, even after all these years, we have to acknowledge that there is still no ideal amino-modifier currently available commercially. Glen Research is now pleased to offer customers a mature version of the amino-modifier C6 phosphoramidite.
Amino-modified oligos are popularly used for microarray manufacturing since DNA chip technology has become such an indispensable tool for life sciences.3,4 Fabrication is based on in situ synthesis on silicon chips or the more accessible approach of post-synthetic immobilization of oligos onto activated surfaces, predominantly glass slides. Companies and oligo synthesis houses use high-throughput processing to prepare libraries of tens of thousands of amino-modified oligos which are then spotted on surfaces to yield microarrays. The spotting process uses either piezo-based dispensers or the more popular pin-based spotters. Procedures have been optimized mainly in terms of immobilization chemistries, with the most popular methods relying on the reaction of amino-modified oligos with surfaces derivatized with moieties such as isothiocyanate or epoxy5 groups. The quality of the product as well as the yield of these amino-modified oligos is vitally important since re-synthesis on this scale is time-consuming and expensive. Clearly, for this kind of high-volume throughput, purification of all these oligos using conventional methods, like HPLC or PAGE, is out of the question - although high-throughput reverse phase (RP) purification in cartridges and plates has proved to be useful.
The most popular amino-modifier used today contains an amine protected by a trifluoroacetyl (TFA) group, e.g., Amino-Modifier C6-TFA (10-1916). This modifier has the advantages of being inexpensive and reliable. However, it has flaws in that it does not allow the use of trityl-on purification techniques or even an estimate of the coupling yield by the trityl cation measurement. Moreover, since the removal of a TFA protecting group happens simultaneously with the deprotection of an oligo, side reactions such as Michael addition of acrylonitrile (formed from elimination of cyanoethyl protecting groups) take place, reducing the yield of the aminated product (Methods to Avoid Inactivation of Primary Amines).
A reagent more suitable for preparation of thousands of amino-modified oligos contains an amine protected with a trityl group, e.g., Amino-Modifier C6 (10-1906). Triphenylmethyl groups (trityls) are a popular family of protecting groups, used in oligonucleotide chemistry for hydroxyl (DMT) and amino (MMT) protection, and removable by mild acidic treatment. Conveniently, trityl cations have large extinction coefficients allowing stepwise coupling yields to be measured easily. Alternatively, due to the hydrophobicity of trityls, separation of the full-length product, still bearing the tritylprotecting group, from capped failure sequences can be carried out, with subsequent acidic removal of the DMT or MMT protection as the final step.
Manufacturing large numbers of oligonucleotides requires cheap and fast purification techniques, ruling out HPLC and PAGE as too expensive and time-consuming. The popular RP cartridge purification method (e.g., PolyPak) does not allow detritylation of the monomethoxytrityl (MMT) group from protected amino-modified oligonucleotides in high yield on the cartridge. The reason for this situation is that acidic cleavage of the N-trityl bond is an equilibrium reaction. On an RP cartridge, where the trityl cation is not physically separated from the amine, the process results in substantial (up to 50%) reattachment of the MMT-group back onto the amine. Subsequent elution of the amino-modified oligonucleotide results in a product that is up to 50% inactive. Interestingly, in addition to this reattachment problem, MMT-amino-modified oligos are quite unstable when stored in aqueous ammonia, gradually losing the MMT group over time.
Recently, we have seen some interest in the dimethoxytrityl equivalent to the amino-modifier C6, presumably in an attempt to overcome some of the problems associated with the MMT version. In our hands, this phosphoramidite is less stable than its MMT cousin and we have concerns about long-term storage. Although the product couples perfectly well in synthesis, there was substantial loss of DMT from the amine during deprotection EVEN at room temperature. Our view is that the DMT-amino-modifier C6 is too labile for routine use but it could prove to be useful in situations requiring very mild removal of the amine protecting group.
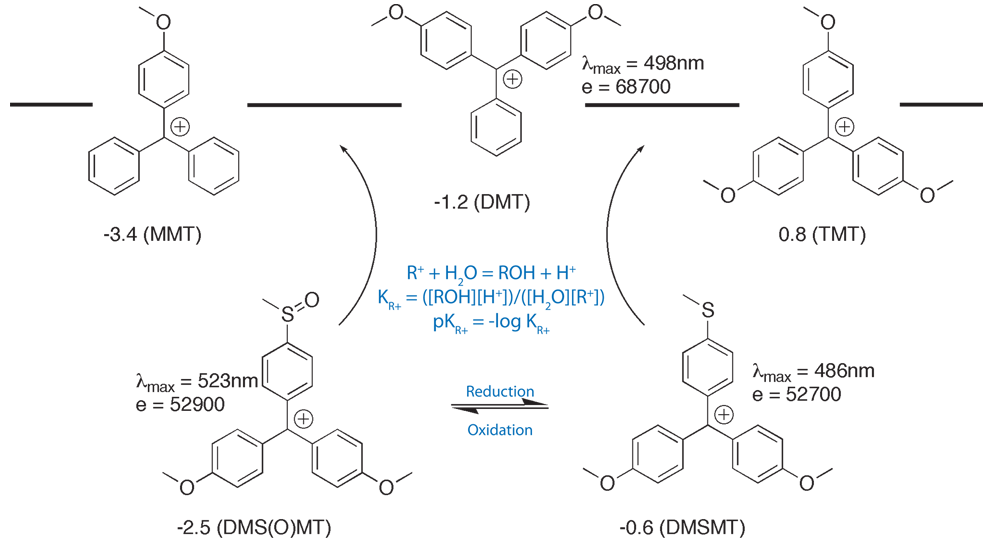
One way around these problems is to use a modified trityl with controlled pKR+, where pKR+ is defined by the following formulae:
R+ + H2O = ROH + H+
KR+ = ([ROH][H+])/([H2O][R+])
pKR+ = -log KR+
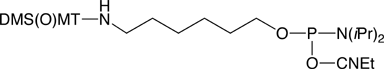
4,4’-Dimethoxy-4’’-thiomethoxytrityl (DMS(O)MT; sulfoxy-form) cation is more stabilized than MMT+, and so the DMS(O)MT-protected amino group is easier to deprotect compared to the MMT-protected one, as shown in Figure 1 on Page 6. The sulfoxy derivative survives conditions of oligonucleotide synthesis and can either be cleaved with standard deblock solution, or left intact for an HPLC purification. At the same time, the DMS(O)MT group is fully compatible with cartridge purification. When detritylation on a cartridge is carried out, the DMS(O)MT+, which is more stable than MMT+, does not reattach itself to an amine. The new aminolink phosphoramidite reagent, 5'-DMS(O)MT-Amino-Modifier C6, utilizing this new trityl based protecting group is shown in Figure 5 (1) on Page 8. The reagent is stable in solution in acetonitrile at room temperature for at least two weeks. UV quantification for release of the new protecting group is possible. Extinction coefficients (L/(mol x cm), shown in Figure 1 on Page 6, were measured in 2% TFA/DCM. In PolyPak detritylation experiments followed by HPLC measurements, the new reagent gave more than 20% improvement in deprotection yields compared to an MMT-protected amino group labeled oligo (4% TFA, 5 min exposure time). A DMS(O)MT-protected amino-modified oligonucleotide was stored in NH4OH/H2O (1:3) at room temperature for 6 days and exhibited no loss of trityl.
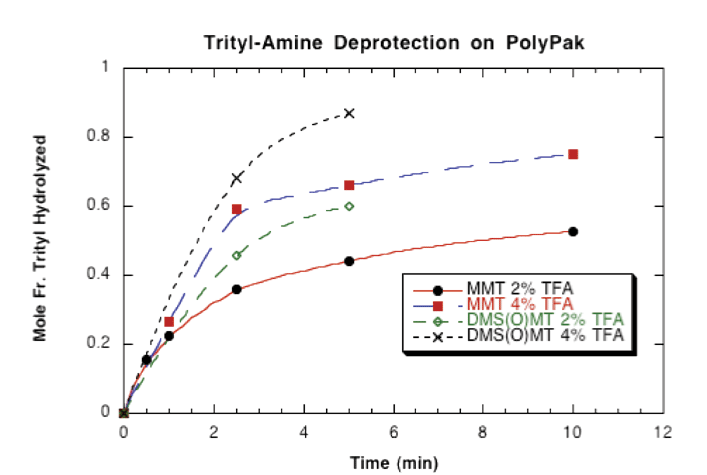
Figure 2 on Page 7 illustrates the rate of deprotection of DMS(O)MT-protected amino-modified oligonucleotides on Poly-Pak cartridges using 2% and 4% aqueous trifluoroacetic acid (TFA). It was found that complete deprotection of DMS(O)MT is achieved after 5 minutes. Under the same conditions, MMT was less than 50% deprotected.
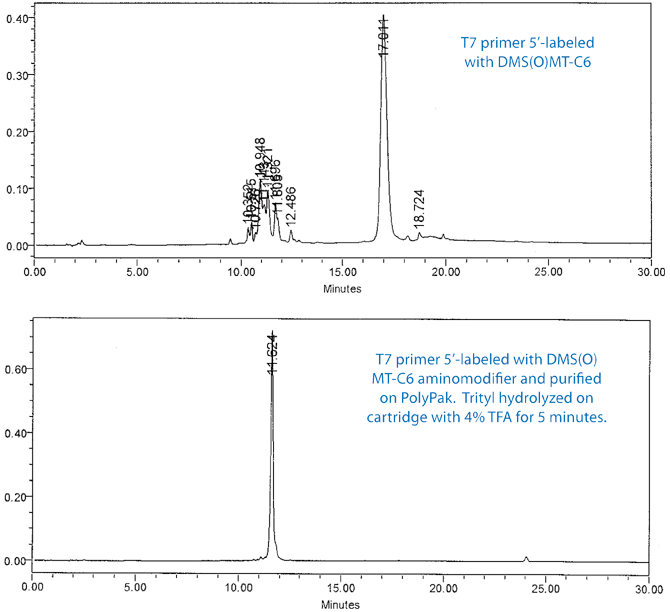
The efficiency of the process is further illustrated in Figure 3 on Page 7. In this experiment, the T7 primer was 5’-labeled with DMS(O)MT-C6 aminomodifier and purified on a PolyPak cartridge. The DS(O)MT trityl was hydrolyzed on the cartridge with 4% TFA for 5 minutes. The upper chromatogram shows the RP HPLC of the DMS(O)MT-on primer and the lower shows the purified amino-modified primer.
Several new applications for dendrimers in the oligonucleotide field have emerged since Glen Research commercialized branching reagents in 1999.6 Examples include multiple fluorescent labeling7, controlled delivery of antisense oligos8, more efficient conjugation of oligos with nano-gold9, and more efficient quenching for molecular beacon applications10. The new trebler described below, now available from Glen Research, possesses features that will assist in some of these applications and thus further expand the array of Glen Research’s building blocks.
The Trebler Phosphoramidite currently available from Glen Research contains a phosphoramidite ‘arm’ that is somewhat crowded by the three adjacent DMT-bearing branches. This leads to increased coupling times (recommended coupling time: 10-15 min) and lower coupling efficiency.
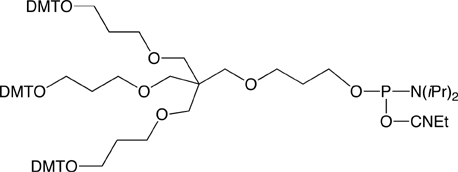
The next generation Trebler (Long Trebler Phosphoramidite) contains an extended phosphoramidite arm, thus markedly reducing the problem of steric hindrance. The new reagent gives higher coupling yields and requires shorter coupling times. Large pore size CPG supports (1000Å and 2000Å) should be selected when using this phosphoramidite. The new trebler phosphoramidite, Long Trebler Phosphoramidite (2), is shown in Figure 5.
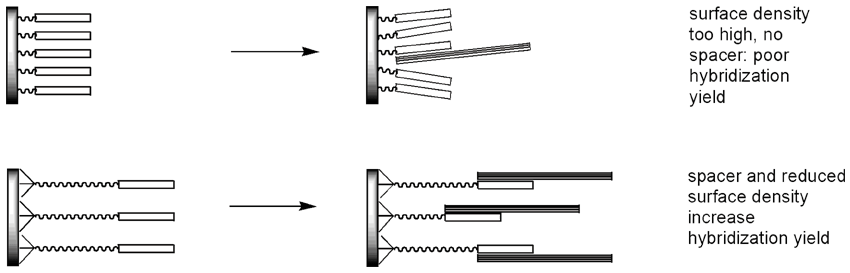
Spatial factors greatly affect the quality of DNA chips. When the probes are too far apart, there would not be enough fluorescence signal to be detected. With probes positioned too close to one another on a surface, there would not be enough room for the target DNA to squeeze through and hybridize - again compromising the quality of the detection. Reducing the surface loading chemically improves the hybridization yield and signal quality, as does linking the oligos to a microarray surface through long spacers. Up to 150 times improvement in hybridization yield compared to oligos directly attached to a surface at high density can be achieved by optimizing spacing and surface density.11
A recently published approach, illustrated in Figure 4, involves a multistep procedure whereby an aminated surface is treated with two-sided bulky structures.12-14 One side of such a structure non-covalently sticks to several reactive sites on the surface, whereas the opposite part of the structure has just one reactive group of its own, thus effectively reducing the surface loading. The method relies on non-covalent binding and requires commercially available glass slides to be additionally modified.
The same result can be achieved in a simpler and more controllable way by employing phosphoramidites available from Glen Research. Upon completion of an oligo synthesis on a large pore CPG (1000Å or 2000Å), the 5’ end can be derivatized with spacer modifiers 9 or 18, followed by a trebler amidite and an amino-modifier amidite. Following standard deprotection, the resulting trebler amino-modified probe will be spotted on to an activated glass surface. The probe will not only immobilize faster courtesy of multiple amino groups, but will also take up more surface area to generate increased spacing, thereby improving the yield in hybridization experiments.
For further technical data on these products please contact: John Thornback at [email protected]
5'-DMS(O)MT-Amino-Modifier C6 (10-1907)
Long Trebler Phosphoramidite (10-1925)