Glen Report 31.15: New Product - 2′-F-5-Me-U-ANA
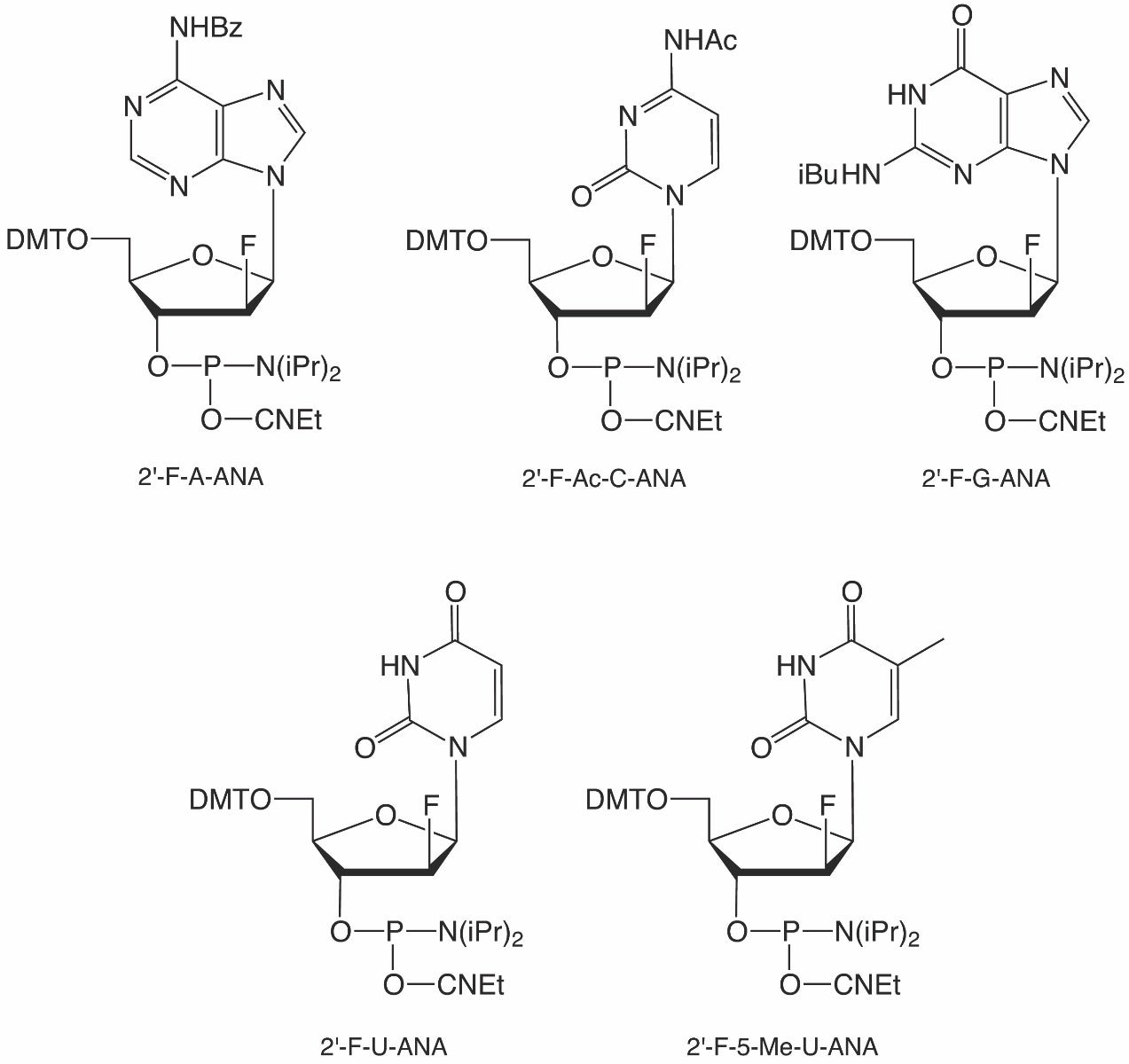
2′-F-arabinonucleic acid (2′-F-ANA) oligonucleotides are cousins to the widely used 2′-F-RNA, with the only difference being an inverted stereocenter at the 2′-carbon. Like 2′-F-RNA, 2′-F-ANA has high base pairing specificity and enhanced hydrolytic and nuclease stability. Glen Research began offering 2′-F-ANA versions of A, C, G and U in 2010 (Figure 1), and since then, this line of products has proven to be popular. Over the years, there have been numerous publications highlighting exciting research using these reagents.
With the goal of enhancing the pharmacological properties of G-quadruplexes, members of the Phan Lab conducted a systematic investigation to understand the effects of several sugar modifications on G-quadruplex structure and stability.1 One of these modifications was 2′-F-ANA. They synthesized a total of twenty singly 2′-F-ANA-modified variants for two types of G-quadruplexes, a (4+0) parallel and a (3+1) hybrid. Using a combination of 1H-NMR, UV absorption and CD, they found that 2′-F-ANA substitutions in anti positions were well tolerated whereas substitutions in syn positions were destabilizing.
In another investigation, Holliger and coworkers investigated several different types of catalysts from non-DNA/RNA backbones such as 2′-F-ANA in the context of the origin of life.2 All their studies involved the use of in vitro selection (SELEX) and previously engineered polymerases.3 Using a self-cleavage strategy, they were able to isolate RNA-cleaving enzymes from an initial random library of 1014 different 2′-F-ANA oligonucleotides after 13-17 rounds of selection. The most active enzyme catalyst was able to cleave RNA in a site- and sequence-specific manner with an observed rate constant of 0.058 min–1. Like DNAzymes and ribozymes, this “FANAzyme” cleaves RNA resulting in the formation of a 5′-hydroxyl group and a 2′,3′- cyclic phosphate. In addition to RNA cleavage, in separate experiments, they were also able to develop FANAzymes that performed the reverse reaction, RNA ligation. The RNA ligase was able to join 3′-imidazolylphosphoryl-RNA to the 5′-OH of another RNA strand, also in a sequence specific manner. Finally, they were able to use a similar selection methodology to also discover a FANAzyme that performed 2′-F-ANA ligation. The ligases exhibited rate constants of 0.0002 and 0.038 min–1, respectively.
Recently, members of the Chaput Lab developed their own RNA-cleaving 2′-F-ANA enzymes as biologically more stable therapeutic candidates.4 Also using SELEX techniques, they were able to isolate superior RNA-cleaving FANAzymes. The most active catalyst was able to cleave RNA in a sequence-specific manner with a maximum rate constant of 0.2 min–1 (Figure 2). Unlike earlier FANAzymes, this catalyst follows Michaelis-Menten kinetics. The researchers confirmed that the enzyme could be re-engineered to target almost any desired RNA sequence by changing the substrate binding domains of the catalyst.
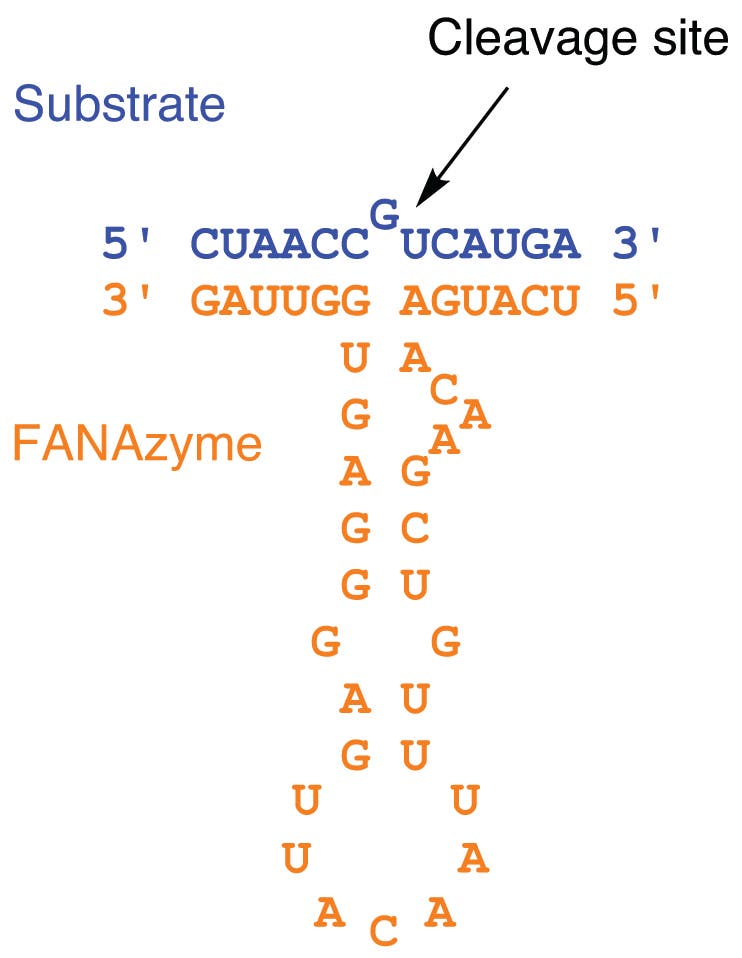
Use of 2′-F-5-Me-U-ANA
Although 2′-F-ANA are epimers of 2′-F-RNA, when 2′-F-ANA hybridizes with DNA, it adopts a more DNA-like B-type helix structure rather than an RNA-like A-type structure. Due to this, it should not come as a surprise that 2′-F-5-Me-U-ANA, which has the extra methyl group similar to thymidine, increases duplex stability relative to 2′-F-U-ANA. To give customers more control in the use of 2′-F-ANA, we are adding 2′-F-5-Me-U-ANA to our catalog (Figure 1). Like our other 2’-F-ANA phosphoramidites, 2′-F-5-Me-U-ANA can be dissolved in acetonitrile and coupled using a coupling time of six minutes using tetrazole as the activator. No changes to standard deprotection methods are required.
References
1. Z. Li, C.J. Lech, and A.T. Phan, Nucleic Acids Research, 2014, 42, 4068-4079.
2. A.I. Taylor, et al., Nature, 2015, 518, 427-30.
3. V.B. Pinheiro, et al., Science, 2012, 336, 341-4.
4. Y. Wang, A.K. Ngor, A. Nikoomanzar, and J.C. Chaput, Nat Commun, 2018, 9, 5067.
Product Information
- Glen Report 31.11: Introducing the New GlenResearch.com
- Glen Report 31.12: 5-Bromo- and 5-Iodo-Pyrimidine Nucleosides; Efficient Oligo to Protein Photo-Cross-linkers (Part 1)
- Glen Report 31.13: New Product - dmf-dG-5’-CPG
- Glen Report 31.14: New Product - Methacrylate C6 Phosphoramidite
- Glen Report 31.15: New Product - 2′-F-5-Me-U-ANA

