Glen Report 7.13: Universal Nucleoside
The design of oligonucleotide primers and probes is frequently made more complicated by the degeneracy of the genetic code or incomplete peptide sequence data. To confront this problem, two strategies are popular but both have significant drawbacks. The use of a mixed base addition (N) at specific points is a logical approach and is achieved either by delivery of equal amounts of monomer on the synthesizer or using the monomers premixed in equivalent amounts. This strategy is more successful the fewer the number of degenerate sites, probably simply due to the smaller number of oligonucleotides formed. (Remember that x degenerate sites lead to 4x oligonucleotides, e.g., 4 fully degenerate sites lead to 256 oligonucleotides of which only one is the desired sequence.) An alternative approach is to substitute 2'-deoxyInosine (dI) at the degenerate sites. Hydrogen bonding of dI to each of the other bases is low but, unfortunately, is not completely equivalent and this factor can cause problems in PCR and sequencing experiments
There clearly exists a need for a new strategy, preferably one that involves a universal base which substitutes each unknown or degenerate site during construction of oligonucleotides. These modified oligonucleotides must still have the ability to generate accurate sequencing data or participate in the amplification of the correct DNA
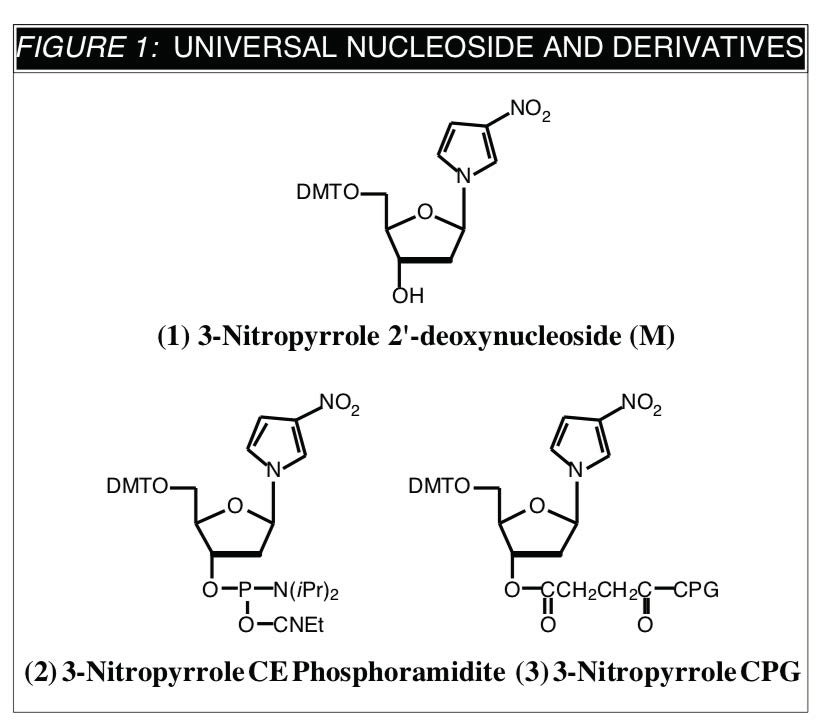
A recent publication1 describes the properties of 3-nitropyrrole 2'-deoxynucleoside (1) (designated M) when used as a universal nucleoside. The authors' strategy was to design and evaluate molecules which would maximize stacking while minimizing hydrogen-bonding interactions. In this way, base-pairing specificity of the target molecule with the 4 normal bases should be minimized while the duplex should be stabilized by the enhanced stacking interactions. For maximum polarity and base stacking, the authors first chose to evaluate 3-substituted pyrrole derivatives as candidates for universal nucleosides based on weakened hydrogen bonding. M was selected from the candidate molecules because of its electronic similarity to p-nitroaniline derivatives which are the smallest known intercalating molecules.
Sequencing
The CE phosphoramidite (2) of 3-nitropyrrole 2'-deoxynucleoside was used to prepare oligonucleotides with several unknown sites substituted with M. The correct sequencing primer in a dideoxy sequencing experiment was compared with the equivalent primers modified at the third position in 4 codons with N (a 256-fold degenerate primer), with dI, and with M. The sequencing ladder obtained using N was unreadable, using dI was only partially readable, but using M was an exact match of the correct oligonucleotide primer. Even more impressively, 17 base oligonucleotide primers with three, six and nine contiguous substitutions of M also generated accurate sequencing data. Both double and single stranded templates were successfully sequenced.
PCR
Promising results were also demonstrated in initial PCR experiments amplifying from total RNA using primers containing three M substitutions. Amplification of the correct product resulted.
Melting Behavior
The ability of an oligonucleotide containing M to function as a sequencing primer would indicate that a duplex is being successfully formed with the complementary strand. Indeed, a normal pattern was observed from the DNA double strand to single strand transition. The Tm values of oligonucleotides containing M-X base pairs (where X=A,C,G or T) all fell within a 3¡ range. In contrast, duplexes containing dI opposite the other bases vary in Tm by as much as 15¡.
Other Potential Applications
As usual, alert and ingenious researchers have already suggested other applications for the use of M: So far, experiments involving ligase chain reaction, in situ hybridization, mutagenesis, motif cloning, and even in restriction fragment length polymorphism (RFLP) have been proposed. We await feedback with interest
As always, Glen Research is dedicated to offering research-oriented products in a timely manner. We are, therefore, happy to offer M as its CE phosphoramidite and the corresponding support (3) under license from the University of Michigan and Purdue University.
Reference
(1) R. Nichols, P.C. Andrews, P. Zhang, and D.E. Bergstrom, Nature, 1994, 369, 492-493.
Product Information
The C5-propynyl-2'-OMe-CE Phosphoramidites and the 3-Nitropyrrole products have been discontinued.

