Glen Report 25.12: Antisense Trimer Phosphoramidites - Update
In our previous Glen Report, GR24.2, we were very pleased to announce the introduction of new Antisense Trimer Phosphoramidites which allow the simultaneous mutation of multiple, distal sites on a gene to be introduced in a facile manner.
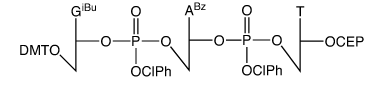
Abbreviated Structure of Trimer Phosphoramidite GAT
However at that time, we did not have any sequencing data that could be used to determine their relative reaction factors (RFs). The RFs are important to compensate for differences between the trimers' relative coupling rates. Trimers that are found to couple slower are given a commensurately larger RF which, in turn, leads to a higher concentration of that trimer phosphoramidite in the trimer mix. The higher concentration of trimer increases the rate of reaction, allowing one to hit the desired target percentage of each codon at the mutation site despite differences in rates of reaction. In the meantime, a value of 1.425 was used, which is the average RF of the well-characterized sense trimers.
In collaboration with Oblique Bio Inc., which specializes in sequence determination, we were able to determine RFs for the new Antisense Trimer Phosphoramidites.
To do so, first a pool of oligos was synthesized with ten incorporations of a twelve-codon trimer mix: 9 antisense trimers and 3 sense trimers for which the RF values are well established. On either side of the block of trimers were fixed flanking sequences for cloning purposes. A primer was annealed to a flanking region and PCR was then used to create a double-stranded pool. These were then cloned into a linearized pGEM vector. After re-circularization and transformation into a DH5 alpha cell line, the clones were plated out onto antibiotic-treated agar plates. The colonies were transferred to liquid media, grown for 18 hours at 37 °C and the plasmids purified by miniprep using the Omega BioTek MagBind purification kit and then sequenced on an ABI 3730xL sequencer. The results are shown in Table 1.
| Trimer | RF | Observed Incorporated (%) | Theoretical Incorporated (%) | Ratio Obs/Theor. | Corrected RFs |
|---|---|---|---|---|---|
| ACC | 1.0 | 9.7 | 8.3 | 1.16 | 0.9 |
| AGA | 1.0 | 6.1 | 8.3 | 0.73 | 1.4 |
| CCA | 1.0 | 7.6 | 8.3 | 0.91 | 1.1 |
| CGG | 1.0 | 10.9 | 8.3 | 1.31 | 0.8 |
| GAT | 1.0 | 5.8 | 8.3 | 0.70 | 1.4 |
| GCA | 1.0 | 8.7 | 8.3 | 1.04 | 1.0 |
| GCG | 1.0 | 13 | 8.3 | 1.56 | 0.6 |
| GGT* | 1.1 | 10.9 | 8.3 | 1.31 | 0.8 |
| GTA | 1.0 | 5.7 | 8.3 | 0.68 | 1.5 |
| TAC* | 1.6 | 8.4 | 8.3 | 1.01 | 1.6 |
| TCT* | 1.3 | 8 | 8.3 | 0.96 | 1.4 |
| TTT | 1.0 | 4.9 | 8.3 | 0.59 | 1.7 |
|
*sense trimer controls |
|||||
With the exception of GGT, the predicted percentages of the sense trimers were quite accurate. The disparity between the observed and the theoretical number of incorporations for a particular trimer may be due to under-sampling or quirks in either the cell line, polymerase, or miniprep, giving a positive or negative selection pressure for that trimer. For this reason, we consider these RF values to be preliminary until we have more sequencing data based upon a variety of amplification and cloning systems. As such, we will refine these values once more sequencing data becomes available and consistent trends are observed. At present, though, we are happy to provide our best estimates for the Antisense Trimer RFs, which are given in Table 2, along with the RFs for the Sense Trimer Phosphoramidites.
| Sense codons (5'->3') | Reaction Factor (RF) | Antisense codons (5'->3') | Reaction Factor (RF) |
|---|---|---|---|
| AAA (Lys) | 1.10 | TTT | 1.70 |
| AAC (Asn) | 1.00 | GTT | 1.90 |
| ACT (Thr) | 1.60 | GGT | 1.10 |
| ATC (Ile) | 1.50 | GAT | 1.40 |
| ATG (Met) | 1.30 | CAT | 1.30 |
| CAG (Gln) | 2.00 | CTG | 1.20 |
| CAT (His) | 1.30 | ATG | 1.30 |
| CCG (Pro) | 1.80 | CGG | 0.80 |
| CGT (Arg) | 1.40 | GCG | 0.60 |
| CTG (Leu) | 1.20 | CAG | 2.00 |
| GAA (Glu) | 1.40 | TTC | 1.30 |
| GAC (Asp) | 1.60 | ATC | 1.50 |
| GCT (Ala) | 1.50 | TGC | 1.50 |
| GGT (Gly) | 1.10 | ACC | 0.90 |
| GTT (Val) | 1.90 | AAC | 1.00 |
| TAC (Tyr) | 1.60 | GTA | 1.50 |
| TCT (Ser) | 1.30 | AGA | 1.40 |
| TGC (Cys) | 1.50 | GCA | 1.00 |
| TGG (Trp) | 1.10 | CCA | 1.10 |
| TTC (Phe) | 1.30 | GAA | 1.40 |
Acknowledgment:
We thank Lance Larka of Oblique Bio Inc. for his assistance with the cloning and sequencing experiments.
Product Information
- Glen Report 25.11: New Product - (5'S)-5',8-Cyclo-dG - DNA Damage and Repair
- Glen Report 25.12: Antisense Trimer Phosphoramidites - Update
- Glen Report 25.13: Technical Brief - Side Reaction of Fluorescein during Deprotection with Methylamine
- Glen Report 25.14: Novel Reagents Utilizing a Serinol Scaffold for Labeling Synthetic Oligonucleotides
- Glen Report 25.15: Technical Brief - Aldehyde and Aminooxy Conjugations
- Glen Report 25.16: Technical Brief - Amino-Modifiers and Summer Shipping

