Glen Report 16.23: Activators, Columns and Plates
Benzylthiotetrazole (BTT)
The first activator described for phosphoramidite chemistry was 1H-tetrazole (1), which has served well over the years. Nevertheless, more potent activators, 5-methylthio-1H-tetrazole (2) and 5-nitrophenyl-1H-tetrazole (3), have popped up but fallen down. More recently, 5-ethylthio-1H-tetrazole (ETT) (4) has gained considerable acceptance as a more acidic activator. ETT has the added advantage of being more soluble in acetonitrile than 1H-tetrazole (up to 0.75M versus 0. 5M solution in acetonitrile). Less acidic but much more nucleophilic is 4,5-dicyanoimidazole (DCI) (5). DCI is even more soluble in acetonitrile (up to 1.2M solution in acetonitrile). ETT and DCI have proved popular for high throughput synthesizers since they do not tend to crystallize and block the fine outlet nozzles.
The renewed interest in RNA synthesis due to the explosion of siRNA technology has led us to evaluate 5-benzylthio-1H-tetrazole (BTT) (6), which was described1 several years ago as an ideal activator for RNA synthesis using TOM-protected RNA phosphoramidites2 and recently3 for TBDMS-protected monomers.
| 1H-tetrazole (1) | 5-methylthio-1H-tetrazole (2) | 5-nitrophenyl-1H-tetrazole (3) |
|---|---|---|
 |
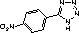 |
|
| 5-ethylthio-1H-tetrazole (4) | 4,5-dicyanoimidazole (5) | Benzylthiotetrazole (BTT) (6) |
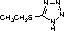 |
 |
Our experiments with BTT have shown that its maximum solubility is about 0.33M in acetonitrile The optimal coupling time for TOM-protected RNA phosphoramidites was found to be 90 seconds and for TBDMS-protected RNA phosphoramidites it was 3 minutes. However, we were concerned that the increased acidity of BTT (pKa 4.08 vs ETT pKa 4.28 vs 1H-tetrazole pKa 4.89)3 could potentially detritylate the monomer during the coupling step leading to a dimer addition. If this process were significant, the full-length oligo would be contaminated with n+1 insertion mutations. However, by examining oligonucleotides by ion-exchange HPLC, we find that n+1 peaks are no more significant using BTT with lower coupling times than ETT with a 6 minute coupling or 1H-tetrazole with a 12 minute coupling.
References
- X. Wu and S. Pitsch, Nucleic Acids Res, 1998, 26, 4315-23.
- S. Pitsch, P.A. Weiss, L. Jenny, A. Stutz, and X.L. Wu, Helv Chim Acta, 2001, 84, 3773-3795.
- R. Welz and S. Muller, Tetrahedron Lett, 2002, 43, 795-797.
Product Information
5-Benzylthio-1H-tetrazole (BTT) Powder (30-3070)
5-Benzylthio-1H-tetrazole (BTT) Solution (30-3170)
Columns and Plates
We are happy to announce the availability of polystyrene columns fully compatible with the Applied Biosystems 3900 synthesizer. Initially, we are offering the regular 4 nucleoside supports, Bz-dA, Bz-dC, dmf-dG and dT. In the near future, we will be adding several of our most popular supports, including RNA supports, for this platform.
We are also adding 96 well plates containing both of our CPG-based universal supports, as well as dT. Universal Support 1000 is suitable for preparing unmodified oligonucleotides. Any regular or modified oligonucleotide, including RNA, can be prepared using Universal Support II. The dT support should find favor among researchers preparing siRNA oligos. These plates are offered initially filled at the 40 nmole level. The support is held tightly in the wells with porous frits, which help to disperse the liquid flow more evenly through the support bed, while avoiding the splashing of support that is virtually inevitable in loosely filled wells.
Product Information
- Glen Report 16.21: Thymine dimers - DNA Lesions Induced By Sunlight CIS-SYN Thymine Dimer Phosphoramidite Now Available
- Glen Report 16.22: EraGen BioSciences Technology (courtesy of EraGen Biosciences)
- Glen Report 16.23: Activators, Columns and Plates
- Glen Report 16.24: Locked Nucleic Acid (LNA™) Phosphoramidites
- Glen Report 16.25: Trimer (Codon) Phosphoramidites Simplify Library Preparation
- Glen Report 16.26: Still More New Products
- Glen Report 16.27: New Epoch Products - 5'-Aldehyde-Modifier C2 Phosphoramidite and Gig Harbor Green™ Phosphoramidite

