Glen Report 13.15: New Monomers: 2,4-Difluorotoluene, Fluorescein Phosphoramidite and Support, 2',3'-Dideoxynucleosides
2,4-Difluorotoluene
Hydrogen bonding between the DNA bases as well as to the amino acid side chains in proteins has been considered to be largely responsible for the specificity of DNA-DNA and DNA-protein interactions. These electrostatic effects are considered to be the primary force behind the exquisite specificity of many polymerases and some nucleases. A wide variety of nucleoside analogues have been examined for their utility in modifying or confusing normal enzymatic functions in medicine and molecular biology. Over the last several years, Eric Kool's group has been working on a base pair which seems to confound conventional logic - there is virtually no hydrogen bonding between the bases.
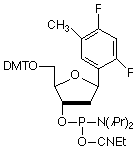
Among several non-polar, aromatic nucleoside analogs that have been studied, 2,4-difluorotoluene (F)1,2 has proved to be remarkable in its ability to act as a non-polar, indeed hydrophobic, mimic of thymidine. F is an identical shape to that of T (Figure 1) and, when substituted for T, is specifically and efficiently replicated3 by Klenow fragment of E. coli DNA polymerase I, a polymerase known to make an error in nucleotide insertion only once in 103 - 105 bases. This is in spite of F's very low capacity for hydrogen bonding. And the corollary works equally well in that F (as triphosphate) is inserted efficiently4 by the polymerase opposite A. Interestingly, melting studies of oligos with F substituting for T indicate4 that the duplex is significantly destabilized. In these studies, F shows little interest in preferring to associate with A over the other bases.
These results would lead to the conclusion that shape selection may be the key factor in base selection by DNA polymerases. Therefore complementarity of shape may be the salient feature in the design of nucleoside analogues designed to probe and modify enzyme interactions. The use of this analogue should allow researchers to probe the level of significance of Watson-Crick hydrogen bonding in any given protein-DNA interaction.
We are delighted to offer the 2,4-difluorotoluene 2'-deoxynucleoside analogue (F) as a phosphoramidite, (1) Figure 3. F is offered under license from the University of Rochester.
References
- B.A. Schweitzer and E.T. Kool, J. Org. Chem., 1994, 59, 7238-7242.
- B.A. Schweitzer and E.T. Kool, J. Am. Chem. Soc., 1995, 117, 1863-1872.
- S. Moran, R.X.-F. Ren, S. Rumney, and E.T. Kool, J. Am. Chem. Soc., 1997, 119, 2056-2057.
- S. Moran, R.X. Ren, and E.T. Kool, Proc Natl Acad Sci U S A, 1997, 94, 10506-11.
Product Information
2,4-Difluorotoluyl Phosphoramidite has been discontinued
6-Fluorescein
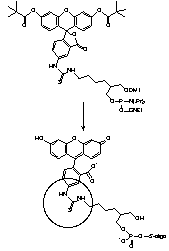
For many years, our most popular fluorescein reagents were Fluorescein Phosphoramidite, (1) Figure 2, and the corresponding Fluorescein CPG. Both of these products are presented on the popular 1,3-diol C7 spacer.1 The fluorescein section of the molecules was derived from fluorescein isothiocyanate (FITC), which, back then, was far and away the most popular reagent for post-synthesis labelling. While these reagents have performed well over the years, recent developments in the design of oligonucleotide probes have made us revisit these products.
So, what's the problem? The isothiocyanate group in FITC, when reacted with amino compound, forms a thiourea linkage, (2) Figure 2. Unfortunately, this linkage is labile during ammonium hydroxide cleavage and deprotection. According to mass spectroscopic evidence from some of our customers, a new species with mass 16 daltons less than expected is also formed after deprotection. It seems clear that this is formed by hydrolysis of the thiourea group to a urea group, (3) Figure 2, (substitution of sulfur by oxygen). A recent publication2 also describes the modification of the thiourea linkage to a guanidinium linkage, (4) Figure 2, by ammonolysis. These results help explain the presence of two or three fluorescein components in the reverse phase HPLC traces of oligos prepared from Fluorescein Phosphoramidite. (The same species are likely present in equivalent 3'-labelled oligos but they do not readily resolve.) It was difficult to explain to a customer that all three peaks in the HPLC represent "the fluorescein oligo". Now add the presence of another fluorophore in doubly-labelled probes and the situation gets really messy.
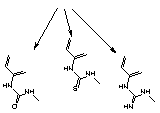
| (3) urea | (2) thiourea | (4) guanidine |
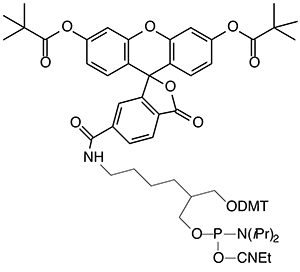
The solution is fairly simple. Use 6-FAM to prepare the fluorescein portion of two new products and they are 6-Fluorescein Phosphoramidite, (2) Figure 3, and 6-Fluorescein CPG, (3) Figure 3. These new products are designed to supplement our existing fluorescein product range and we would envisage that the original two products will be phased out as demand drops off.

References
- P.S. Nelson, M. Kent, and S. Muthini, Nucleic Acids Res., 1992, 20, 6253-6259.
- I. Dubey, G. Pratviel, and B. Meunier, Bioconjug Chem, 1998, 9, 627-32.
Product Information
6-Fluorescein Phosphoramidite (10-1964)
3'-(6-Fluorescein) CPG (20-2964)
2',3'-Dideoxynucleosides
Many of customers have expressed an interest in preparing oligos with a 2',3'-dideoxynucleoside at the 3'-terminus. Unfortunately, the 2',3'-dideoxynucleoside supports are very difficult to prepare and only ddC is currently available from us. An alternative approach to the preparation of these oligos is to synthesize in the 5'-3' sense, using 5'-supports and phosphoramidites, with the 2',3'-dideoxynucleoside 5'-phosphoramidite being added to the 3'-terminus in the last cycle. Unfortunately, since the 2',3'-dideoxynucleoside 5'-phosphoramidite has no DMT group, this approach is incompatible with DMT-on purification techniques. Also, since failure sequences contain a 3'-OH group, it is imperative that they be removed from the product by ion-exchange HPLC or gel electrophoresis. The structures of the 2',3'-dideoxynucleoside monomers are shown, (4) - (7), Figure 3.
Product Information
- Glen Report 13.11: C-5 Propynes, Cytofectin and Other Product Updates
- Glen Report 13.12: A Rapid Method for Atomic Mutagenesis of Nucleic Acids
- Glen Report 13.13: Trityl Group in the 3rd Millenium: New Perspectives for Oligonucleotide Chemistry and Beyond
- Glen Report 13.14: Polyplex Synthesizer from Genemachines
- Glen Report 13.15: New Monomers: 2,4-Difluorotoluene, Fluorescein Phosphoramidite and Support, 2',3'-Dideoxynucleosides
- Glen Report 13.16: Product Update - Which 5'-Amino-Modifier?