Glen Report 28.22: New Product - TIPS-5-Ethynyl-dU-CE Phosphoramidite
First introduced in 2002, Copper(I)-catalyzed azide-alkyne cycloaddition (CuAAC) is a robust and versatile Click Chemistry methodology commonly characterized by high yields, simple conditions, mild reagents, innocuous by-products, and simple purification.1-3 Subsequent developments by Carell and Seela with copper-stabilizing ligands that protect DNA from copper-induced damage and alkyne-bearing modifications for oligonucleotide synthesis have produced a versatile method that is fully compatible with oligonucleotide labelling.4-8
In 2010, Glen Research partnered with baseclick GmbH to offer alkyne-bearing phosphoramidites, azides, and ancillary reagents for labelling oligonucleotides using CuAAC Click conjugations. Through an updated agreement with baseclick we are pleased to continue to offer and expand our list of these versatile reagents for labelling of oligonucleotides for research and development use.
Previous work incorporating 5-Ethynyl-dU-CE Phosphoramidite (1) in oligonucleotide synthesis showed that this modified nucleoside worked well, albeit with some restrictions. Unlike the C8-Alkyne-dT-CE Phosphoramidite (2), 5-ethynyl-dU is subject to base-catalyzed hydration during cleavage and deprotection, especially when using a strong base or heat.9 Hydration of an ethynyl group forms a methyl ketone which subsequently blocks potential click reactions. Mild deprotection conditions are necessary to prevent this side reaction. Acid catalyzed reactions are also possible for 5-ethynyl-dU although we did not observe any degradation in our short test oligos.
In this article, we introduce TIPS-5-Ethynyl-dU-CE Phosphoramidite (3). This protected alkyne offers broader compatibility with oligonucleotide synthesis and deprotection. Protecting the 5-ethynyl group with a triisopropylsilyl (TIPS) protecting group prevented acid or base catalyzed hydration during oligonucleotide synthesis and workup.9 A quick treatment with TBAF removed the TIPS protecting group. A simple ethanol precipitation or Glen Gel-Pak™ desalting step produced very pure oligos suitable for CuAAC click conjugations.
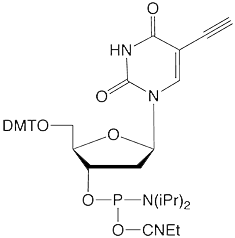
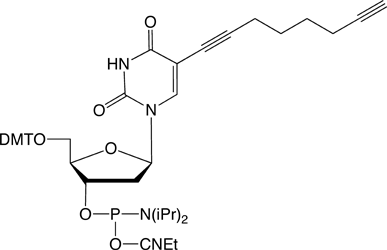
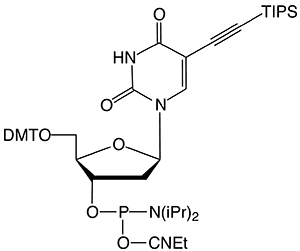
Figure 1: 5-Ethynyl-dU, its TIPS Protected Analogue and C8-Alkyne-dT |
||
 |
 |
 |
(1) 5-Ethynyl-dU-CE Phosphoramidite |
(2) C8-Alkyne-dT-CE Phosphoramidite |
(3) TIPS-5-Ethynyl-dU-CE Phosphoramidite |
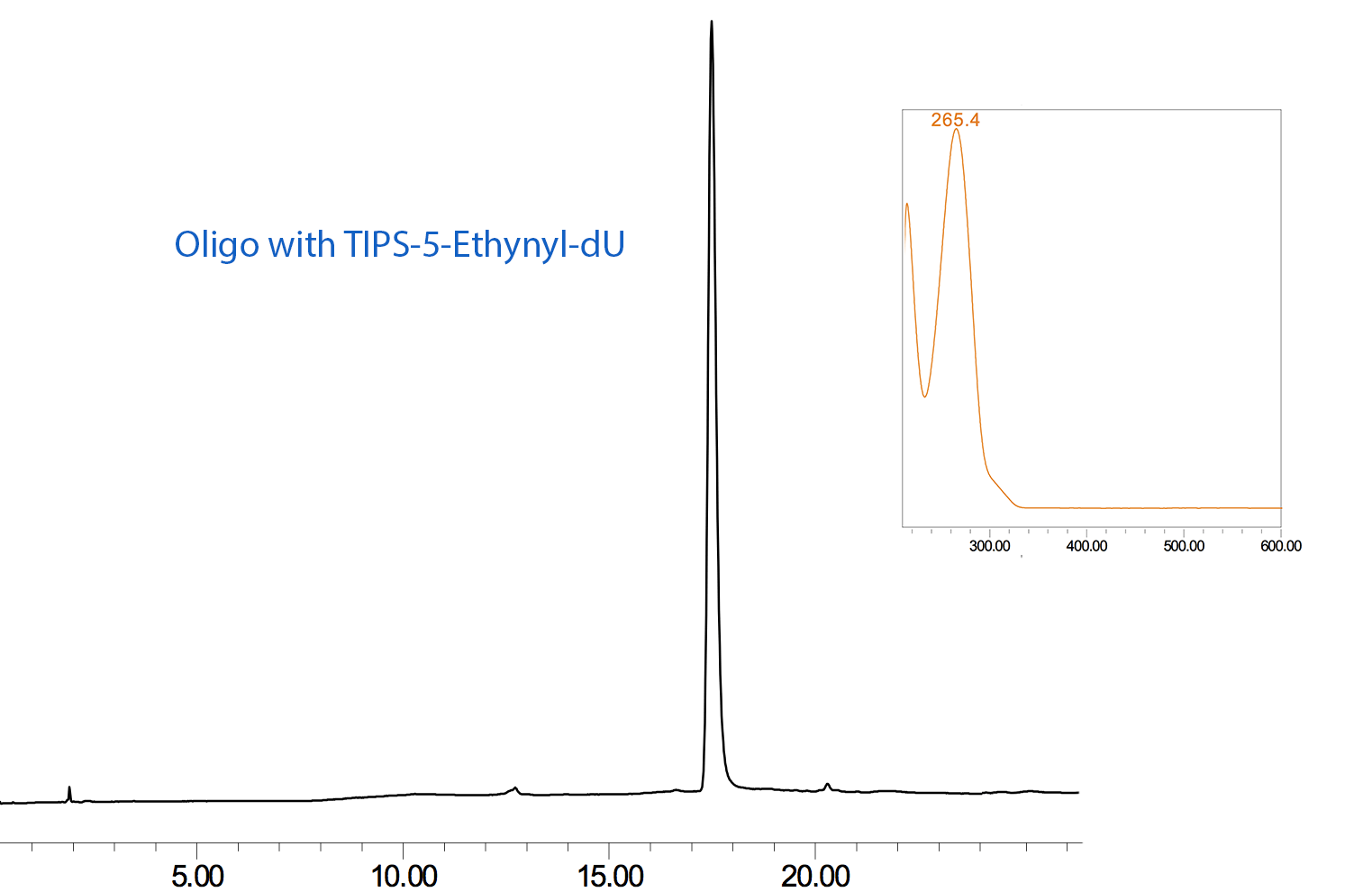
Figure 2: RP HPLC Analysis of 12-Mer Oligo Before and After Click Conjugation with HEX


Product Information
TIPS-5-Ethynyl-dU Phosphoramidite requires a 3 minute coupling time. Oligonucleotide cleavage and deprotection can be carried out according to the requirements of the nucleobases. After deprotection, the oligonucleotide is dried. A variety of protocols are then available for removing the TIPS protecting group but we found the following simple protocol worked very well.
Protocol for TIPS removal
- Dissolve the oligonucleotide in 0.4mL anhydrous DMF.
- Transfer the solution to a plastic centrifuge tube.
- Add 0.1mL tetrabutylammonium fluoride (TBAF) to the tube.
- Deprotect at 45°C for 15 minutes.
- Quench the reaction with 0.5mL 2M triethylammonium acetate (TEAA).
- Desalt the oligonucleotide on a Glen Gel-Pak cartridge, or equivalent, and filter the product solution if necessary.
Product Highlights
5-Ethynyl-dU Phosphoramidite compatibility
✔ ammonium hydroxide deprotection at room temperature
✗ not compatible with ammonium hydroxide AND heat
✔ potassium carbonate deprotection
✗ not compatible with any AMA deprotection
TIPS-5-Ethynyl-dU Phosphoramidite compatibility
✔ ammonium hydroxide at 55°C for 17 hours
✔ ammonium hydroxide deprotection at room temperature
✔ potassium carbonate deprotection
✔ AMA deprotection at 65°C for 10 minutes
✔ AMA deprotection at room temperature for 2 hours
Previous Glen Reports have highlighted strategies and applications for Click conjugation.10 Some of the diverse strategies include solid and aqueous-phase conjugations, small and large-scale conjugations, and multiple click reporters. Applications include crosslinking, ligation, spin labels, and photo-crosslinking. We offer a diverse set of reagents for modifying oligonucleotides, including terminal modifiers, internal nucleosidic modifiers, non-nucleosidic modifiers, and post synthesis modifiers.
References
1. V.V. Rostovtsev, Green, L.G., Fokin, V.V. and Sharpless, K.B., Angew Chem Int Ed, 2002, 41, 2596-2599.
2. C.W. Tornøe, C. Christensen, and M. Meldal, The Journal of Organic Chemistry, 2002, 67, 3057-3064.
3. H.C. Kolb, M.G. Finn, and K.B. Sharpless, Angewandte Chemie International Edition, 2001, 40, 2004-2021.
4. P.M.E. Gramlich, S. Warncke, J. Gierlich, and T. Carell, Angewandte Chemie-International Edition, 2008, 47, 3442-3444.
5. P.M.E. Gramlich, C.T. Wirges, A. Manetto, and T. Carell, Angewandte Chemie-International Edition, 2008, 47, 8350-8358.
6. F. Seela, and V.R. Sirivolu, Helvetica Chimica Acta, 2007, 90, 535-552.
7. F. Seela, and V.R. Sirivolu, Organic & Biomolecular Chemistry, 2008, 6, 1674-1687.
8. F. Seela, V.R. Sirivolu, and P. Chittepu, Bioconjugate Chemistry, 2008, 19, 211-224.
9. S.A. Ingale, H. Mei, P. Leonard, and F. Seela, The Journal of Organic Chemistry, 2013, 78, 11271-82.
10. The Glen Report, 2014, 26.1, 7-8. https://www.glenresearch.com/reports/gr26-15
Product Information
5-Ethynyl-dU-CE Phosphoramidite (10-1554)
C8-Alkyne-dT-CE Phosphoramidite(10-1540)
TIPS-5-Ethynyl-dU-CE Phosphoramidite (10-1555)
- Glen Report 28.21: Versatile Applications of the Copper(I)-Catalyzed Click Chemistry
- Glen Report 28.22: New Product - TIPS-5-Ethynyl-dU-CE Phosphoramidite
- Glen Report 28.23: Technical Brief – Fluorescein Labelling – Considering the Options
- Glen Report 28.24: New Products – AquaPhluor® 593 5’-Phosphoramidite and CPG
- Glen Report 28.25: New Product: Glen-Pak™ DNA 3mg 384-Well Plate
- Glen Report 28.26: Transient Cyanoethylation - An Unexpected Route to Chemical Bleaching of a Fluorophore

