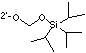Glen Report 11.21: Advances in RNA Synthesis and Structural Analysis
RNA Synthesis
In March 1991, we reviewed1 the state of RNA synthesis and concluded that the monomers we offered at that time, with 2'-O-t-butyldimethylsilyl (TBDMS) and labile base protecting groups, were close to being optimal. In the intervening years, we have continued to supply these monomers and some incremental improvements have been made, including the use of ethylthiotetrazole2,3 as a more efficient activator, acetyl protected C monomer along with methylamine2,4 for deprotection, and triethylamine trihydrofluoride5 for desilylation. With these modifications, RNA synthesis is currently very healthy but the process is still not as dependable or as routine as DNA synthesis.
However, two recent innovations promise to push RNA synthesis to new levels, as well as introduce two new acronyms, ACE and TOM, to the field.
The key to a new synthesis strategy6 is the use of 2'-O-bis(2-acetoxyethoxy)methyl (ACE) orthoester protection which requires substitution of DMT with silyl ethers for 5' protection. Although substantial cycle and reagent changes are required in this strategy, its great advantage is the stability of the 2'-O-protected RNA which can be purified and stored. The 2'-protection is then efficiently removed by incubation in aqueous buffers. This strategy holds distinct promise and we look forward to the further optimization and commercialization of this chemistry.
Although the TBDMS group has served well for 2' protection over many years, a deceptively simple change to 2'-O-triisopropylsilyloxymethyl (TOM)7 protection offers very significant advantages which are detailed in an account from Xeragon AG beginning on Page 2. Glen Research has contracted with Xeragon to supply TOM RNA monomers exclusively worldwide.
NAIM
In addition to these recent developments in the field of RNA synthesis, a powerful and elegant new technique is now available for probing the structure and function of RNA molecules. Nucleotide Analog Interference Mapping8 (NAIM) allows one to probe the effect of substituting an analog for a particular nucleotide in all positions within an RNA molecule simultaneously. The NAIM process is reviewed in detail by Scott Strobel beginning on Page 6.
NAIM is a radical departure from the conventional means of determining the contribution of a functional group to the activity of an RNA. Current techniques have relied upon exhaustive analysis of a series of substituted RNAs prepared by either chemical9,10 or semi-synthetic11 methods. Rather than using such a brute force approach, NAIM utilizes a combinatorial strategy in which a population of substituted RNAs is generated and analyzed. By using a-thiotriphosphate nucleotide analogs, the sites affected by the substitution, exhibiting either inhibition or enhancement of activity, are determined without anything more esoteric than a sequencing gel. At present, NAIM has been used to refine the structural basis for the activity of Tetrahymena group I intron12,13,14, but may be readily extended in any system where a selection process between the active and inactive species is possible including catalysis, protein or ligand interactions, and folding for RNA and even DNA molecules.
References
- Glen Research Report, 1991, 4, 1-4.
- F. Wincott, et al., Nucleic Acids Res., 1995, 23, 2677-2684.
- B. Sproat, et al., Nucleosides and Nucleotides, 1995, 14, 255-273.
- M.P. Reddy, N.B. Hanna, and F. Farooqui, Nucleosides and Nucleotides, 1997, 16, 1589-1598.
- D. Gasparutto, et al., Nucleic Acids Res., 1992, 20, 5159-5166.
- S.A. Scaringe, F.E. Wincott, and M.H. Caruthers, J Am Chem Soc, 1998, 120, 11820-11821.
- X. Wu and S. Pitsch, Nucleic Acids Res., 1998, 26, 4315-23.
- S.A. Strobel and K. Shetty, Proc Natl Acad Sci U S A, 1997, 94, 2903-8.
- W.A. Pieken, D.B. Olsen, F. Benseler, H. Aurup, and F. Eckstein, Science, 1991, 253, 314-7.
- A.M. Pyle and T.R. Cech, Nature, 1991, 350, 628-31.
- M.J. Moore and P.A. Sharp, Science, 1992, 256, 992-7.
- S. Basu, et al., Nat Struct Biol, 1998, 5, 986-92.
- S.A. Strobel, L. Ortoleva-Donnelly, S.P. Ryder, J.H. Cate, and E. Moncoeur, Nat Struct Biol, 1998, 5, 60-6.
- L. Ortoleva-Donnelly, A.A. Szewczak, R.R. Gutell, and S.A. Strobel, RNA, 1998, 4, 498-519.
Product Information
Natural RNA Phosphoramidites and Supports
The a-thiotriphosphates have been discontinued.
- Glen Report 11.21: Advances in RNA Synthesis and Structural Analysis
- Glen Report 11.22: TOM-Protecting-Group™ - A Major Improvement in RNA Synthesis
- Glen Report 11.23: New Columns for Expedite and LV Applications
- Glen Report 11.24: Nucleotide Analogs for Interference Mapping of RNA
- Glen Report 11.25: Alternatives to Expedite Monomers
- Glen Report 11.26: More Novel Monomers