Glen Report 20.11: An Unnatural Base Pair System for the Expansion of Genetic Information
Expansion of the genetic alphabet by unnatural base pair systems is an attractive future biotechnology.1 DNA fragments containing unnatural base pairs function as templates for the site-specific incorporation of extra components into nucleic acids and proteins by biosyntheses, according to the Central Dogma of molecular biology - replication, transcription, and translation. Nucleic acids and proteins with extra components at specific positions could exhibit novel or increased functionality, which would be useful for both basic research and applied sciences. Several researchers have been actively developing such unnatural base pairs, some of which display unique abilities in duplex DNA and in nucleic acid and protein biosyntheses.2-5 Glen Research is already a source for nucleoside derivatives related to unnatural base pairs, such as dP, dK, 5-Me-isodC, and isodG. Now we are adding to our lineup the amidites of dDs, dPa, and ds, the elements for a new unnatural base pair system.
Ds--Pa Base Pair
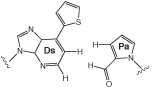
Base Pair Relies on Hydrophobicity and Shape
The unnatural base pair between 7-(2-thienyl)-imidazo[4,5-b]pyridine (Ds) and pyrrole-2-carbaldehyde (Pa) is formed by specific hydrophobic shape complementation. The shape of the Ds-Pa pair is different from those of the natural A-T and G-C pairs, but the Ds-Pa pair works together with the natural pairs in in vitro replication and transcription (Figures 1 and 2 on Page 2).6 The Ds-Pa pair complementarily functions as a template base in DNA fragments for exonuclease-proficient (exo+) DNA polymerases, such as the Klenow fragment and Vent DNA polymerase for replication, as well as T7 RNA polymerase for transcription. In replication by the Klenow fragment (exo+), the triphosphate substrates of the unnatural bases, dDsTP and dPaTP, are incorporated into DNA opposite the respective Pa and Ds bases in the template strands under the conventional conditions. In addition, the DNA fragments containing the Ds-Pa pair can be amplified by PCR using Vent DNA polymerase. This PCR amplification requires6 modified substrates, γ-amidotriphosphates of Ds and A, instead of their corresponding triphosphate substrates, to increase the fidelity. Under specific conditions, the total mutation rate of the Ds-Pa site in DNA fragments after 10 PCR cycles was estimated to be ~1%.
The Ds-Pa pair also complementarily mediates6 the site-specific incorporation of Ds, Pa, and modified Pa bases into RNA by conventional T7 transcription. For example, the ribonucleoside triphosphate of Pa (PaTP) or biotin-linked Pa (Biotin-PaTP) can be incorporated at desired positions in RNA molecules opposite Ds in DNA templates by T7 RNA polymerase. Biotin-linked Pa is used for the immobilization of RNA molecules on avidin supports and also as a chemiluminescent marker, using streptavidin coupled to alkaline phosphatase, to detect RNA molecules and their interactions with other molecules.
s--Pa Base Pair
Pa also functions as a template base for incorporating another unnatural base (Figures 1 and 2 on Page 2), 2-amino-6-(2-thienyl)purine (s), into RNA.6 As described below, the s base acts as a unique fluorescent base analog in DNA and RNA fragments.
s--y Base Pair
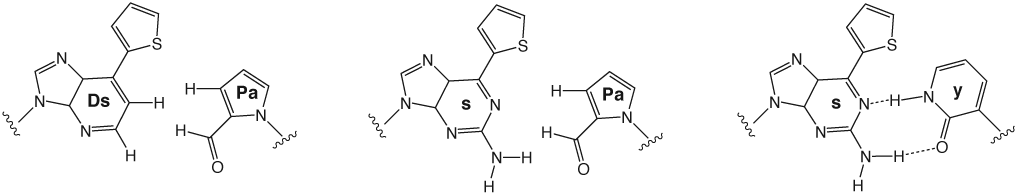
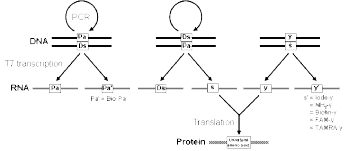
The s base was originally developed as a pairing partner of 2-oxopyridine (y), and the s-y pair is compatible with both transcription and translation (Figures 1 and 2).7 The substrate of y (yTP) for transcription is site-specifically incorporated into RNA opposite s in DNA templates by T7 RNA polymerase. Modified y bases, such as iodo-, aminoalkyl-, biotin-linked, and fluorophore-linked y bases, can also be incorporated into RNA by specific transcription using s-containing DNA templates.8 Iodo-y is a photosensitive component that is capable of cross-linking upon irradiation at 312 nm,9 and aminoalkyl-y is useful for post-transcriptional modification at the aminoalkyl-y incorporation site in RNA molecules.10 Biotin- or fluorophore-linked y was used in an RNA aptamer to facilitate the detection of the interaction between the aptamer and its target molecules.10,11
RNA molecules containing y at a specific position can be used as mRNA for the expansion of the genetic code, in which novel codons containing y are assigned to unnatural amino acids. The combination of this specific transcription with in vitro translation systems enables protein synthesis incorporating extra amino acids with functional groups of interest. In the translation system, tRNA with an s-containing anticodon is required to interact with the corresponding y-containing codon in mRNA. The preparation of the s-containing tRNA molecules is accomplished by T7 transcription using Pa-containing DNA templates and sTP by mediating the s-Pa pairing, in which Pa is the more efficient template base than y for the s incorporation into RNA.12
Fluorescence Properties of s Base
Interestingly, the s base itself is strongly fluorescent (excitation: 299 or 352 nm; emission: 434 nm; quantum yield: 41%).12,13 The fluorescent intensity of s in oligonucleotides decreases upon stacking with neighboring bases, and thus the fluorescent profiles reflect the local structure at the labeling site of the oligonucleotides. In addition, fluorescence resonance energy transfer (FRET) is observed between a low-fluorescence state of s and a high-fluorescence state of another fluorophore-linked y, such as 5-carboxyfluorescein-linked y (FAM-y), in oligonucleotides.12 Using these abilities of s, researchers can design a variety of ingenious structural analyses of DNA and RNA molecules.
Oligonucleotide Synthesis
DNA fragments containing Ds or Pa are prepared by DNA chemical synthesis using the dDs- or dPa-amidite by the usual phosphoramidite methods.
DNA fragments containing s are prepared by DNA chemical synthesis using the ds-amidite, in which the amino group is protected with a phenoxyacetyl group. We, therefore, recommend the use of phenoxyacetic anhydride (Pac2O) in Cap A for the capping process during DNA synthesis. This is because the use of the usual acetic anhydride capping reagent exchanges some of the phenoxyacetyl protecting groups on s with the acetyl groups. RNA fragments containing s are prepared by T7 transcription using sTP and Pa-containing DNA templates.13
Initially, Glen Research will supply researchers with dDs-, dPa-, and ds-amidites (Figure 3 on Page 3) for DNA chemical synthesis. Glen Research is indebted to TagCyx Biotechnologies for incorporating these phosphoramidites into our Agreement. TagCyx Biotechnologies has licenses from RIKEN and The University of Tokyo (for Ds and Pa) and from Japan Science and Technology Agency (for s).
We are indebted to Professor Hirao for providing us with text for this article.
It is our pleasure to inform our readers that some triphosphates mentioned in this article will shortly be available from Glen Research. Specifically, sTP and Biotin-PaTP will be added to our catalog in the near future. Some others will follow at a later date. Please check our web site for further updates. Also, subscribe to our e-mail newsletter by sending an e-mail to custserv@glenres.com to learn immediately about the release of these triphosphates.
References
- S.A. Benner, P. Burgstaller, T.R. Battersby, and S. Jurczyk, Did the RNA world exploit an expanded genetic alphabet? In The RNA World, Gesteland. R.F.; Cech T.R.; Atkins J.F., Eds. Cold Spring Harbor Laboratory Press: 1999; pp 163-181.
- J.D. Bain, C. Switzer, A.R. Chamberlin, and S.A. Benner, Nature, 1992, 356, 537-9.
- E.T. Kool, Curr Opin Chem Biol, 2000, 4, 602-608.
- A.A. Henry, and F.E. Romesberg, Curr Opin Chem Biol, 2003, 7, 727-33.
- I. Hirao, Curr Opin Chem Biol, 2006, 10, 622-7.
- I. Hirao, et al., Nat Methods, 2006, 3, 729-35.
- I. Hirao, et al., Nat Biotechnol, 2002, 20, 177-82.
- I. Hirao, Biotechniques, 2006, 40, 711-717.
- M. Kimoto, et al., Chem Biol, 2004, 11, 47-55.
- R. Kawai, et al., J Am Chem Soc, 2005, 127, 17286-95.
- K. Moriyama, M. Kimoto, T. Mitsui, S. Yokoyama, and I. Hirao, Nucl. Acids Res., 2005, 33, e129.
- T.Mitsui, M.Kimoto, R.Kawai, S.Yokoyama, and I.Hirao, Tetrahedron, 2007, 35, 5360-5369.
- M. Kimoto, et al., Nucl. Acids Res., 2007, 35, 5360-5369.
Intellectual Property
This product is covered by patents or patents pending owned by TagCyx Biotechnologies. Purchase of this product includes a limited license to use this product solely for research. This license specifically excludes: (a) therapeutic or diagnostic applications (including products or services that incorporate this product), (b) any in vivo toxicity/safety study in support of an investigational new drug application (or foreign counterpart), (c) resale, or (d) gene functionalization activities (including products or services that incorporate data derived from gene functionalization activities) if such activities have commercial application. All of the above require a separate license from TagCyx Biotechnologies. Neither this product nor any product created through its use may be used in human clinical trials.
Please also see: Site-specific incorporation of functional components into RNA by transcription using unnatural base pair systems
Product Information
dDs-CE Phosphoramidite, Pac-ds-CE Phosphoramidite and dPa-CE Phosphoramidite have been discontinued
- Glen Report 20.11: An Unnatural Base Pair System for the Expansion of Genetic Information
- Glen Report 20.12: Thiophosphoramidites and Their Use in Synthesizing Oligonucleotide Phosphorodithioate Linkages
- Glen Report 20.13: Technical Brief - Purification of 6-FAM Labelled oligos using Glen-Pak™ Cartridges
- Glen Report 20.14: More Click Chemistry
- Glen Report 20.15: Technical Brief - New application of 5-Me-iso-dC and iso-dG
- Glen Report 20.16: Technical Brief - New application for 5’-OMe-dT phosphoramidite
- Glen Report 20.17: New Product - Deuterated 2’-Deoxyguanosine Phosphoramidite
- Glen Report 20.18: New Products - Solid CPR II / Formylindole – Aldehyde Modifier
- Glen Report 20.19:New Product - 5'-Cholesteryl-TEG Phosphoramidite
- Glen Report 20.110: Improving Universal Support II for Oligonucleotide Synthesis
- Glen Report 20.111: Differences Between Universal Support II and III
- Glen Report 20.112: Technical Brief - Phage Display, Artificial Antibodies and Trimer Phosphoramidites

