Jeffrey R. Vieregg and Niles A. Pierce
Department of Bioengineering,
Department of Computing and Mathematical Sciences,
California Institute of Technology,
Pasadena, California 91125
Nucleic acid probes are used for diverse applications in vitro, in situ, and in vivo. In any setting, their power is limited by imperfect selectivity (binding of undesired targets) and incomplete affinity (binding is reversible and not all desired targets are bound). These difficulties are fundamental, stemming from reliance on base pairing to provide both selectivity and affinity. Chemical modifications to base or backbone moieties within the probe can partially address these challenges, but come with their own set of drawbacks. Shielded covalent (SC) probes eliminate the longstanding tradeoff between selectivity and durable target capture, achieving selectivity via programmable base pairing and molecular conformation change, and durable target capture via activatable covalent crosslinking (Figure 1a)1.
The SC probe concept is suitable for use with diverse crosslinker chemistries, including 3-cyanovinylcarbazole (CNVK), a photoactive nucleoside analog developed by Yoshimura et al. (Figure 1b)2, 3. We found that the CNVK phosphoramidite coupled with high yield using the same conditions as unmodified nucleosides and is compatible with traditional ammonium hydroxide deprotection at room temperature (see Reference 1 for details). We observed crosslinking yields above 90% after UV-A activation with either high-intensity LED or low-cost fluorescent lamps. The resulting crosslinks are stable against either thermal or chemical denaturation and, if desired, can be efficiently reversed with UV-B light. The only sequence requirement for CNVK crosslinking is a single opposite-strand pyrimidine, allowing a wide choice of target sequences.
In pure and mixed samples, SC probes covalently capture complementary DNA or RNA oligonucleotide targets and reject two-nucleotide mismatched targets with near-quantitative yields at room temperature, achieving discrimination ratios of 2–3 orders of magnitude. Semi-quantitative studies with full-length mRNA targets demonstrate selective covalent capture comparable to that for RNA oligo targets. Single-nucleotide DNA or RNA mismatches, including nearly-isoenergetic RNA wobble pairs, can be efficiently rejected with discrimination ratios of 1–2 orders of magnitude. Covalent capture yields appear consistent with the thermodynamics of probe/target hybridization, facilitating rational probe design1.
SC probes require only one modified nucleoside (the crosslinker) and can incorporate DNA, RNA or modified bases as desired. In contrast to existing probe chemistries, SC probes achieve the high sequence selectivity of a structured probe, yet durably retain their targets even under denaturing conditions. This previously incompatible combination of properties suggests diverse applications based on selective and stable binding of nucleic acid targets under conditions where base-pairing is disrupted (e.g., by stringent washes in vitro or in situ, or by enzymes in vivo).
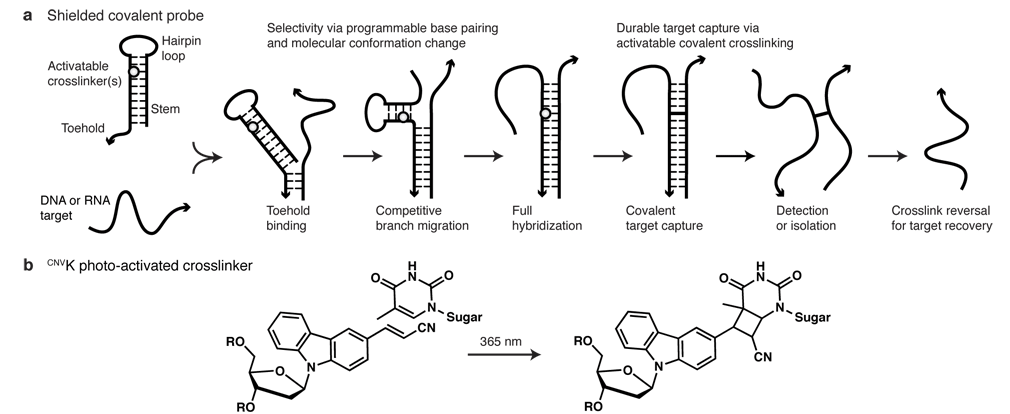
Adapted with permission from Vieregg et al., J. Am. Chem. Soc., 135:9691-9699, 2013. Copyright 2013 American Chemical Society.
CNVK is covered by US Patent 8,481,714 and other patents worldwide assigned to the Japan Advanced Institute of Science and Technology (JAIST). At the time of writing (December 2013), CNVK phosphoramidite was not available commercially.