First came the genetic code mediated by the four regular DNA bases. But beyond genetics lies the recently explored space of epigenetics. Our little corner of this space is dedicated to oligonucleotide synthesis and, for us, epigenetics predominantly means methylated cytosine variants and their incorporation into oligonucleotides. Figure 1 shows the structures of dC and its epigenetically significant variants. The C5 methylated version of dC, 5-Me-dC, is oxidized stepwise to 5-hydroxymethyl-dC (5-hm-dC), 5-formyl-dC (5-f-dC) and 5-carboxy-dC (5-ca-dC) (Figure 1).

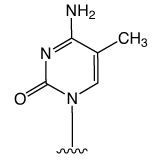
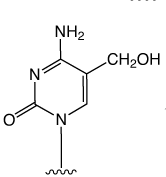
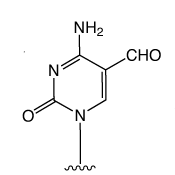

In 2009, Glen Research introduced the 5-hm-dC phosphoramidite (1) and that product has proved to be very useful even though it requires very forcing conditions (ammonium hydroxide at 75 °C for 17 hours) for complete deprotection. In 2011, we published an article by Markus Mueller and Thomas Carell of the Ludwig-Maximilians University in Munich, which detailed the processes involved in cytosine methylation and demethylation. At that time, we introduced a second monomer for hm-dC (2), and monomers for f-dC (3) and ca-dC (4). The structures of these monomers are shown in Figure 2. Using these monomers, researchers were able to prepare oligonucleotides containing one or several of these individual bases but could they produce an oligo containing all of these bases? Unfortunately, the answer was – no – but why?
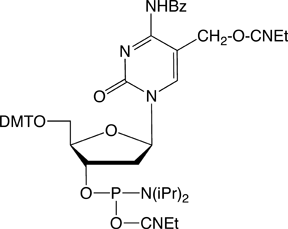
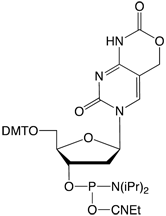

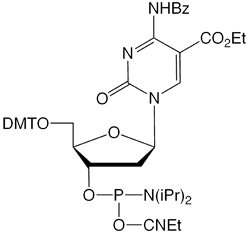

Figure 2: Phosphoramidite MonomersFirst, ca-dC (4) is not compatible with deprotection using ammonium hydroxide or AMA which would lead to a mixture of the desired carboxylic acid and incorrect amides. So, an oligo containing all four of these methylated analogues has to be deprotected using sodium hydroxide in aqueous methanol to be compatible with ca-dC. Fortunately, hm-dC II (2) is also compatible with sodium hydroxide deprotection, as indeed is f-dC (3).However, the structure of the f-dC monomer (3) requires that the fully deprotected oligo be treated with 50mM aqueous sodium periodate to generate the formyl group (Figure 3). This treatment has been reported to be incompatible with hm-dC (1).1

Clearly a new f-dC structure is required. The previous effort to achieve this goal was to prepare the 5-formyl-dC monomer with no protection on the formyl group. Although this product almost solved the problem, the reactivity of the unprotected formyl group precluded making the more complex oligonucleotides that researchers needed.After preparing and testing a variety of candidates, the Carell group was able to design a monomer for f-dC which seems to meet all of the requirements to prepare an oligo containing all of the methylated variants.1 The structure of the f-dC III monomer (5) is shown in Figure 2.The aldehyde is protected as an acetal group and the exocyclic amino group is protected with the 4-methoxy-benzoyl group. The choice of the acetal protecting group is critical since the group must be easily removed while surviving the conditions of oligonucleotide synthesis. It was found that the acetal formed from propane-1,3-diol offered the optimal characteristics. It was also found that the regular N4-benzoyl protecting group was too labile in f-dC and its partial loss during oligonucleotide synthesis led to some chain branching. So, the more resistant 4-methoxy-benzoyl group was chosen in its place.Using the f-dC III monomer, an oligo containing all four of these methylated bases can be simply prepared. The synthesis conditions are normal and the deprotection is carried out using 0.4M sodium hydroxide in methanol/water 4:1 (v/v) for 17 hours at room temperature. (Please note that this deprotection scheme is not compatible with the use of dmf-protected dG in the oligo. However, iBu-dG is fully deprotected under these conditions.) After deprotection with sodium hydroxide, the oligo must not be isolated by simple evaporation. Instead, the sodium hydroxide can be neutralized using a suitable buffer or the oligo can be ethanol precipitated.To remove the acetal protecting group, as shown in Figure 4, the oligo is treated with acetic acid/water 80:20 (v/v) for 6 hours at 20 °C. It should be noted that both time and temperature are critical for complete removal of the acetal protecting group with absence of side reactions.

These highly modified oligos can be simply purified by Glen-Pak purification. The normal procedure can be followed with the exception that the removal of the 5'-DMT group with 2% aqueous trifluoroacetic acid should be omitted. The DMT group is then removed during the acetic acid/water procedure described above for removal of the acetal protecting groups.The work of the Carell group described here is a superb illustration of the use of protecting groups in oligonucleotide synthesis. We are happy to offer these products for sale and to acknowledge that bringing this set of monomers to market would not have been possible without their talent and help.Reference
| Building Block*/ Deprotection Conditions | NH4OH or AMA | NaOH (not compatible with dmf-dG) | K2CO3 |
|---|---|---|---|
| 5‑Hydroxymethyl‑dC (1) | Ammonium hydroxide at 75° C for 17 hours. | Not compatible | Not compatible |
| 5‑Hydroxymethyl‑dC II (2) | Yields urea derivatives – possibility to introduce different ureas post-synthetically by deprotection with suitable amine | 0.4 M NaOH in methanol/water 4:1, 25°C, 17 h | 0.05M K2CO3 in anhydrous methanol, 25°C, 17 h using UltraMild conditions |
| 5‑Formyl‑dC (3) | Standard deprotection followed by periodate oxidation, see Figure 3 | Standard deprotection followed by periodate oxidation, see Figure 3 | Standard deprotection followed by periodate oxidation, see Figure 3 |
| 5‑Formyl‑dC III (5) | Conc. NH4OH, 25°C, 17 h followed by acetal removal, see Figure 4 | 0.4 M NaOH in methanol/water 4:1, 25°C, 17 h followed by acetal removal, see Figure 4 | 4-Methoxy-benzoyl protecting group is not compatible with UltraMild deprotection conditions |
| 5-Carboxy-dC (4) | Yields amides as well as desired carboxylic acid | 0.4 M NaOH in methanol/water 4:1, 25°C, 17 h | Not compatible |
| *See Figure 2 for structures | |||
5-Hydroxymethyl-dC II-CE Phosphoramidite (10-1510)
5-Formyl dC III CE Phosphoramidite (10-1564)
5-Carboxy-dC-CE Phosphoramidite (10-1066)