Researchers have long been searching for fluorescent nucleosides to probe DNA structure. Fluorescent nucleoside bases would also be useful in the analysis of DNA-protein interactions.
Etheno-A (1) and etheno-C (2)1 are two readily accessible fluorescent structures but these molecules are both non-hybridizing. Indeed, etheno-dA is a mutagenic nucleoside formed by the action of vinyl chloride on DNA. Other notable fluorescent base analogues are the pteridine nucleoside analogues actively being investigated by Pfleiderer, Hawkins and co-workers. Although very challenging from a synthetic standpoint, these analogues hold great promise because of their substantial fluorescence, as well as their potential to hybridize specifically with natural bases. The most promising analogue described to date2 is the adenosine analogue (3) but guanosine and other analogues have also been intensively investigated.3-5 2-Aminopurine (4) is a mildly fluorescent base, which has found significant use in probing DNA structures. It is especially useful in that it is capable of hybridizing with T (Figure 4) in Watson-Crick geometry.6
Two years ago, in our constant search for nucleosides with interesting fluorescence properties, we introduced furano-dT (7). We had no reason to believe that this molecule would exhibit any base pairing properties and we expected it to act like etheno-dA in that regard. Still, there is always a need for a novel fluorescent nucleoside to aid in DNA structural analysis, so we launched it as an interesting alternative fluorescent base. Before long, it became apparent that there was more to this product than we initially observed. We were aware that this base was not very stable to deprotection conditions, although normal ammonium hydroxide treatment seemed to cause no problems. However, other techniques using potassium carbonate or sodium hydroxide caused decomposition of the base, thereby losing its fluorescent properties. The next indication we had that there were problems inherent with this structure came when we examined mass spectrometric data on oligonucleotides containing multiple additions of furano-dT. Consistently, the molecular weights were 1 Dalton less per addition of furano-dT than expected. When we checked melting behavior of oligonucleotides containing furano-dT, the answer quickly became clear. Melting curves generated using oligos containing "furano-dT" with complementary strands containing A, C, G or T opposite the "furano-dT" sites clearly revealed the solution. "Furano-dT" did not hydrogen bond to A, C, or T, but formed a perfect base pair with G. Mass spec and melting data were trying to tell us that furano-dT had been transformed in the ammonium hydroxide cleavage and deprotection to the equivalent amino compound, which we have chosen to call pyrrolo-dC (6) (Figure 1).
A literature search quickly generated further evidence that furano-dT is not stable to ammonia, which converts it to the equivalent pyrrolo-pyrimidine, pyrrolo-dC. Interestingly, researchers at Epoch7 who synthesized a pyrrolo-dC analogue were examining potential base pairs for selectively binding complementary (SBC) oligo-nucleotides. They did not note the fluorescent properties of the pyrrolo-pyrimidine ring system. However, they did observe that their version of this ring system, different from pyrrolo-dC only in lacking the methyl group on the pyrrolo ring, formed a mismatch with G and was quite unstable to deprotection conditions.
In the meantime, one group had already applied "furano-dT" to their research endeavors. We were able to alert them to the transformation to pyrrolo-dC in ammonia. In their publication8, they made use of the fact that the fluorescence intensity of pyrrolo-dC is sensitive to its environment and can be used to provide direct information on local melting of the DNA. This example makes use of the fact that the fluorescence is quenched in duplex DNA relative to its fluorescence in single-stranded DNA.
In response to the instability of furano-dT to base, we have discontinued the phosphoramidite monomer (7) from our catalog (although it is still available to any researchers who are continuing research started with it). We now introduce Pyrrolo-dC CE Phosphoramidite (8) and look forward to this monomer having a long, stable existence as a Glen Research specialty, while allowing researchers to probe DNA structure and activity using a fluorescent base that forms a perfect match to G on the complementary strand.
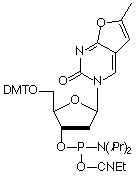
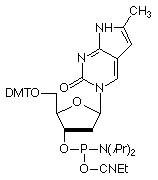
Figure 3 shows the normalized excitation and emission spectra of Pyrrolo-dC in a single-stranded oligonucleotide. The fluorescence is, of course, partially quenched in double-stranded oligonucleotides.
In the meantime, we continue to carry out experiments designed to investigate the properties of pyrrolo-dC and we will report on these in detail in the coming months.
| λmaxima: 265 and 347 nm | hybidization state QY |
|---|---|
| E260: 4000 L/mol cm single-stranded | 0.07 |
| E347: 3700 L/mol cm double-stranded | 0.02 |